HP1α mediates defective heterochromatin repair and accelerates senescence in Zmpste24-deficient cells.
Liu, J; Yin, X; Liu, B; Zheng, H; Zhou, G; Gong, L; Li, M; Li, X; Wang, Y; Hu, J; Krishnan, V; Zhou, Z; Wang, Z
Cell cycle (Georgetown, Tex.)
13
1237-47
2014
Mostrar resumen
Heterochromatin protein 1 (HP1) interacts with various proteins, including lamins, to play versatile functions within nuclei, such as chromatin remodeling and DNA repair. Accumulation of prelamin A leads to misshapen nuclei, heterochromatin disorganization, genomic instability, and premature aging in Zmpste24-null mice. Here, we investigated the effects of prelamin A on HP1α homeostasis, subcellular distribution, phosphorylation, and their contribution to accelerated senescence in mouse embryonic fibroblasts (MEFs) derived from Zmpste24(-/-) mice. The results showed that the level of HP1α was significantly increased in Zmpste24(-/-) cells. Although prelamin A interacted with HP1α in a manner similar to lamin A, HP1α associated with the nuclease-resistant nuclear matrix fraction was remarkably increased in Zmpste24(-/-) MEFs compared with that in wild-type littermate controls. In wild-type cells, HP1α was phosphorylated at Thr50, and the phosphorylation was maximized around 30 min, gradually dispersed 2 h after DNA damage induced by camptothecin. However, the peak of HP1α phosphorylation was significantly compromised and appeared until 2 h, which is correlated with the delayed maximal formation of γ-H2AX foci in Zmpste24(-/-) MEFs. Furthermore, knocking down HP1α by siRNA alleviated the delayed DNA damage response and accelerated senescence in Zmpste24(-/-) MEFs, evidenced by the rescue of the delayed γ-H2AX foci formation, downregulation of p16, and reduction of senescence-associated β-galactosidase activity. Taken together, these findings establish a functional link between prelamin A, HP1α, chromatin remodeling, DNA repair, and early senescence in Zmpste24-deficient mice, suggesting a potential therapeutic strategy for laminopathy-based premature aging via the intervention of HP1α. | | | 24584199
 |
DNA methylation analysis of the macrosatellite repeat associated with FSHD muscular dystrophy at single nucleotide level.
Huichalaf, C; Micheloni, S; Ferri, G; Caccia, R; Gabellini, D
PloS one
9
e115278
2014
Mostrar resumen
Facioscapulohumeral muscular dystrophy (FSHD) is one of the most common inherited diseases of the skeletal muscle. It is characterized by asymmetric muscle weakness and variable penetrance. FSHD is linked to a reduction in copy number of the D4Z4 3.3 kb macrosatellite repeat, located in 4q35. This causes the epigenetic de-repression of FSHD candidate genes leading to disease. Nevertheless, the molecular mechanism responsible for silencing of FSHD candidate genes in healthy subjects is not fully understood. While a role for DNA methylation has been suggested, so far there is limited information regarding the methylation status of the 325 CpGs contained in each D4Z4 unit. Using a human/rodent monochromosomal hybrid cell line containing a single human chromosome 4, we performed an in depth analysis of DNA methylation for the majority of the CpGs inside D4Z4 at single nucleotide level. We found that D4Z4 is not uniformly methylated and that the level of DNA methylation does not correlate with the density of CpG dinucleotides. Moreover, in several D4Z4 regions characterized by near complete methylation, we found specific unmethylated CpGs. These elements are enriched in transcription factor binding sites that could be involved in muscle-specific D4Z4 activity. Our approach also detected differential methylation among different D4Z4 units, suggesting that the D4Z4 array is a mosaic of euchromatic and heterochromatic domains. Finally, we found that DNA methylation and histone de-acetylation are required to maintain FSHD candidate genes repressed. Taken together, our data underscore new players involved in the epigenetic regulation of the FSHD locus that could be targeted for therapeutic purposes. | | | 25545674
 |
Proteomic and genomic approaches reveal critical functions of H3K9 methylation and heterochromatin protein-1γ in reprogramming to pluripotency.
Sridharan, R; Gonzales-Cope, M; Chronis, C; Bonora, G; McKee, R; Huang, C; Patel, S; Lopez, D; Mishra, N; Pellegrini, M; Carey, M; Garcia, BA; Plath, K
Nature cell biology
15
872-82
2013
Mostrar resumen
Reprogramming of somatic cells into induced pluripotent stem cells (iPSCs) involves a marked reorganization of chromatin. To identify post-translational histone modifications that change in global abundance during this process, we have applied a quantitative mass-spectrometry-based approach. We found that iPSCs, compared with both the starting fibroblasts and a late reprogramming intermediate (pre-iPSCs), are enriched for histone modifications associated with active chromatin, and depleted for marks of transcriptional elongation and a subset of repressive modifications including H3K9me2/me3. Dissecting the contribution of H3K9 methylation to reprogramming, we show that the H3K9 methyltransferases Ehmt1, Ehmt2 and Setdb1 regulate global H3K9me2/me3 levels and that their depletion increases iPSC formation from both fibroblasts and pre-iPSCs. Similarly, we find that inhibition of heterochromatin protein-1γ (Cbx3), a protein known to recognize H3K9 methylation, enhances reprogramming. Genome-wide location analysis revealed that Cbx3 predominantly binds active genes in both pre-iPSCs and pluripotent cells but with a strikingly different distribution: in pre-iPSCs, but not in embryonic stem cells, Cbx3 associates with active transcriptional start sites, suggesting a developmentally regulated role for Cbx3 in transcriptional activation. Despite largely non-overlapping functions and the predominant association of Cbx3 with active transcription, the H3K9 methyltransferases and Cbx3 both inhibit reprogramming by repressing the pluripotency factor Nanog. Together, our findings demonstrate that Cbx3 and H3K9 methylation restrict late reprogramming events, and suggest that a marked change in global chromatin character constitutes an epigenetic roadblock for reprogramming. | | | 23748610
 |
A NF-κB-dependent dual promoter-enhancer initiates the lipopolysaccharide-mediated transcriptional activation of the chicken lysozyme in macrophages.
Witham, J; Ouboussad, L; Lefevre, PF
PloS one
8
e59389
2013
Mostrar resumen
The transcriptional activation of the chicken lysozyme gene (cLys) by lipopolysaccharide (LPS) in macrophages is dependent on transcription of a LPS-Inducible Non-Coding RNA (LINoCR) triggering eviction of the CCCTC-binding factor (CTCF) from a negative regulatory element upstream of the lysozyme transcription start site. LINoCR is transcribed from a promoter originally characterized as a hormone response enhancer in the oviduct. Herein, we report the characterization of this cis-regulatory element (CRE). In activated macrophages, a 60 bp region bound by NF-κB, AP1 and C/EBPβ controls this CRE, which is strictly dependent on NF-κB binding for its activity in luciferase assays. Moreover, the serine/threonine kinase IKKα, known to be recruited by NF-κB to NF-κB-dependent genes is found at the CRE and within the transcribing regions of both cLys and LINoCR. Such repartition suggests a simultaneous promoter and enhancer activity of this CRE, initiating cLys transcriptional activation and driving CTCF eviction. This recruitment was transient despite persistence of both cLys transcription and NF-κB binding to the CRE. Finally, comparing cLys with other LPS-inducible genes indicates that IKKα detection within transcribing regions can be correlated with the presence of the elongating form of RNA polymerase II or concentrated in the 3' end of the gene. | | | 23533622
 |
Autophagy-dependent senescence in response to DNA damage and chronic apoptotic stress.
Singh, K; Matsuyama, S; Drazba, JA; Almasan, A
Autophagy
8
236-51
2011
Mostrar resumen
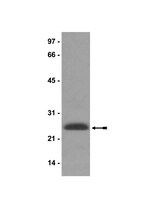
Autophagy regulates cell survival and cell death upon various cellular stresses, yet the molecular signaling events involved are not well defined. Here, we established the function of a proteolytic Cyclin E fragment (p18-CycE) in DNA damage-induced autophagy, apoptosis, and senescence. p18-CycE was identified in hematopoietic cells undergoing DNA damage-induced apoptosis. In epithelial cells exposed to DNA damage, chronic but not transient expression of p18-CycE leads to higher turnover of LC3 I/II and increased emergence of autophagosomes and autolysosomes. Levels of p18-CycE, which was generated by proteolytic cleavage of endogenous Cyclin E, were greatly increased by chloroquine and correlated with LC 3II conversion. Preventing p18-CycE genesis blocked conversion of LC3 I to LC3 II. Upon DNA damage, cytoplasmic ataxia-telangiectasia-mutated (ATM) was phosphorylated in p18-CycE-expressing cells resulting in sustained activation of the adenosine-mono-phosphate-dependent kinase (AMPK). These lead to sustained activation of mammalian autophagy-initiating kinase ULK1, which was abrogated upon inhibiting ATM and AMPK phosphorylation. Moreover, p18-CycE was degraded via autophagy followed by induction of senescence. Both autophagy and senescence were prevented by inhibiting autophagy, which leads to increased apoptosis in p18-CycE-expressing cells by stabilizing p18-CycE expression. Senescence was further associated with cytoplasmic co-localization and degradation of p18-CycE and Ku70. In brief, chronic p18-CycE expression-induced autophagy leads to clearance of p18-CycE following DNA damage and induction of senescence. Autophagy inhibition stabilized the cytoplasmic p18-CycE-Ku70 complex leading to apoptosis. Thus, our findings define how chronic apoptotic stress and DNA damage initiate autophagy and regulate cell survival through senescence and/or apoptosis. | Western Blotting | | 22240589
 |
Heterochromatin protein 1 gamma and IκB kinase alpha interdependence during tumour necrosis factor gene transcription elongation in activated macrophages.
Thorne, JL; Ouboussad, L; Lefevre, PF
Nucleic acids research
40
7676-89
2011
Mostrar resumen
IκB kinase α (IKKα) is part of the cytoplasmic IKK complex regulating nuclear factor-κB (NF-κB) release and translocation into the nucleus in response to pro-inflammatory signals. IKKα can also be recruited directly to the promoter of NF-κB-dependent genes by NF-κB where it phosphorylates histone H3 at serine 10, triggering recruitment of the bromodomain-containing protein 4 and the positive transcription elongation factor b. Herein, we report that IKKα travels with the elongating form of ribonucleic acid polymerase II together with heterochromatin protein 1 gamma (HP1γ) at NF-κB-dependent genes in activated macrophages. IKKα binds to and phosphorylates HP1γ, which in turn controls IKKα binding to chromatin and phosphorylation of the histone variant H3.3 at serine 31 within transcribing regions. Downstream of transcription end sites, IKKα accumulates with its inhibitor the CUE-domain containing protein 2, suggesting a link between IKKα inactivation and transcription termination. | | Mouse | 22649058
 |
CBX3 regulates efficient RNA processing genome-wide.
Smallwood, A; Hon, GC; Jin, F; Henry, RE; Espinosa, JM; Ren, B
Genome research
22
1426-36
2011
Mostrar resumen
CBX5, CBX1, and CBX3 (HP1α, β, and γ, respectively) play an evolutionarily conserved role in the formation and maintenance of heterochromatin. In addition, CBX5, CBX1, and CBX3 may also participate in transcriptional regulation of genes. Recently, CBX3 binding to the bodies of a subset of genes has been observed in human and murine cells. However, the generality of this phenomenon and the role CBX3 may play in this context are unknown. Genome-wide localization analysis reveals CBX3 binding at genic regions, which strongly correlates with gene activity across multiple cell types. Depletion of CBX3 resulted in down-regulation of a subset of target genes. Loss of CBX3 binding leads to a more dramatic accumulation of unspliced nascent transcripts. In addition, we observed defective recruitment of splicing factors, including SNRNP70, to CBX3 target genes. Collectively, our data suggest a role for CBX3 in aiding in efficient cotranscriptional RNA processing. | | | 22684280
 |
Suppression and recovery of BRCA1-mediated transcription by HP1γ via modulation of promoter occupancy.
Choi, JD; Park, MA; Lee, JS
Nucleic acids research
40
11321-38
2011
Mostrar resumen
Heterochromatin protein 1γ (HP1γ) is a chromatin protein involved in gene silencing. Herein, we show that HP1γ interacts with breast cancer type 1 susceptibility protein (BRCA1) and regulates BRCA1-mediated transcription via modulation of promoter occupancy and histone modification. We used several HP1γ mutants and small interfering RNAs for histone methyltransferases to show that BRCA1-HP1γ interaction, but not methylated histone binding, is important in HP1γ repression of BRCA1-mediated transcription. Time-lapse studies on promoter association and histone methylation after DNA damage revealed that HP1γ accumulates at the promoter before DNA damage, but BRCA1 is recruited at the promoter after the damage while promoter-resident HP1γ is disassembled. Importantly, HP1γ assembly recovers after release from the damage in a BRCA1-HP1γ interaction-dependent manner and targets SUV39H1. HP1γ/SUV39H1 restoration at the promoter results in BRCA1 disassembly and histone methylation, after which transcription repression resumes. We propose that through interaction with BRCA1, HP1γ is guided to the BRCA1 target promoter during recovery and functions in the activation-repression switch and recovery from BRCA1-mediated transcription in response to DNA damage. | | | 23074186
 |
Dynamics and memory of heterochromatin in living cells.
Hathaway, NA; Bell, O; Hodges, C; Miller, EL; Neel, DS; Crabtree, GR
Cell
149
1447-60
2011
Mostrar resumen
Posttranslational histone modifications are important for gene regulation, yet the mode of propagation and the contribution to heritable gene expression states remains controversial. To address these questions, we developed a chromatin in vivo assay (CiA) system employing chemically induced proximity to initiate and terminate chromatin modifications in living cells. We selectively recruited HP1α to induce H3K9me3-dependent gene silencing and describe the kinetics and extent of chromatin modifications at the Oct4 locus in fibroblasts and pluripotent cells. H3K9me3 propagated symmetrically and continuously at average rates of ~0.18 nucleosomes/hr to produce domains of up to 10 kb. After removal of the HP1α stimulus, heterochromatic domains were heritably transmitted, undiminished through multiple cell generations. Our data enabled quantitative modeling of reaction kinetics, which revealed that dynamic competition between histone marking and turnover, determines the boundaries and stability of H3K9me3 domains. This framework predicts the steady-state dynamics and spatial features of the majority of euchromatic H3K9me3 domains over the genome. | | | 22704655
 |
Reversal of heterochromatic silencing of quiescent herpes simplex virus type 1 by ICP0.
Ferenczy, MW; DeLuca, NA
Journal of virology
85
3424-35
2010
Mostrar resumen
Persisting latent herpes simplex virus genomes are to some degree found in a heterochromatic state, and this contributes to reduced gene expression resulting in quiescence. We used a relatively long-term quiescent infection model in human fibroblasts, followed by provision of ICP0 in trans, to determine the effects of ICP0 on the viral chromatin state as gene expression is reactivated. Expression of ICP0, even at low levels, results in a reduction of higher-order chromatin structure and heterochromatin on quiescent viral genomes, and this effect precedes an increase in transcription. Concurrent with transcriptional activation, high levels of ICP0 expression result in the reduction of the heterochromatin mark trimethylated H3K9, removal of histones H3 and H4 from the quiescent genome, and hyperacetylation of the remaining histones. In contrast, low levels of ICP0 did not appreciably change the levels of histones on the viral genome. These results indicate that ICP0 activity ultimately affects chromatin structure of quiescent genomes at multiple levels, including higher-order chromatin structure, histone modifications, and histone association. Additionally, the level of ICP0 expression affected its ability to change chromatin structure but not to reactivate gene expression. While these observations suggest that some of the effects on chromatin structure are possibly not direct, they also suggest that ICP0 exerts its effects through multiple mechanisms. | | | 21191021
 |