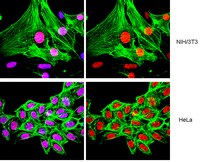
Phosphorylation and arginine methylation mark histone H2A prior to deposition during Xenopus laevis development.
Wang, WL; Anderson, LC; Nicklay, JJ; Chen, H; Gamble, MJ; Shabanowitz, J; Hunt, DF; Shechter, D
Epigenetics & chromatin
7
22
2014
Zobrazit abstrakt
Stored, soluble histones in eggs are essential for early development, in particular during the maternally controlled early cell cycles in the absence of transcription. Histone post-translational modifications (PTMs) direct and regulate chromatin-templated transactions, so understanding the nature and function of pre-deposition maternal histones is essential to deciphering mechanisms of regulation of development, chromatin assembly, and transcription. Little is known about histone H2A pre-deposition modifications nor known about the transitions that occur upon the onset of zygotic control of the cell cycle and transcription at the mid-blastula transition (MBT).We isolated histones from staged Xenopus laevis oocytes, eggs, embryos, and assembled pronuclei to identify changes in histone H2A modifications prior to deposition and in chromatin. Soluble and chromatin-bound histones from eggs and embryos demonstrated distinct patterns of maternal and zygotic H2A PTMs, with significant pre-deposition quantities of S1ph and R3me1, and R3me2s. We observed the first functional distinction between H2A and H4 S1 phosphorylation, as we showed that H2A and H2A.X-F (also known as H2A.X.3) serine 1 (S1) is phosphorylated concomitant with germinal vesicle breakdown (GVBD) while H4 serine 1 phosphorylation occurs post-MBT. In egg extract H2A/H4 S1 phosphorylation is independent of the cell cycle, chromatin assembly, and DNA replication. H2AS1ph is highly enriched on blastula chromatin during repression of zygotic gene expression while H4S1ph is correlated with the beginning of maternal gene expression and the lengthening of the cell cycle, consistent with distinct biological roles for H2A and H4 S1 phosphorylation. We isolated soluble H2A and H2A.X-F from the egg and chromatin-bound in pronuclei and analyzed them by mass spectrometry analysis to quantitatively determine abundances of S1ph and R3 methylation. We show that H2A and H4 S1ph, R3me1 and R3me2s are enriched on nucleosomes containing both active and repressive histone PTMs in human A549 cells and Xenopus embryos.Significantly, we demonstrated that H2A phosphorylation and H4 arginine methylation form a new class of bona fide pre-deposition modifications in the vertebrate embryo. We show that S1ph and R3me containing chromatin domains are not correlated with H3 regulatory PTMs, suggesting a unique role for phosphorylation and arginine methylation. | Immunoblotting (Western) | 25302076
 |
Analysis of histones in Xenopus laevis. I. A distinct index of enriched variants and modifications exists in each cell type and is remodeled during developmental transitions.
Shechter, D; Nicklay, JJ; Chitta, RK; Shabanowitz, J; Hunt, DF; Allis, CD
The Journal of biological chemistry
284
1064-74
2009
Zobrazit abstrakt
Histone proteins contain epigenetic information that is encoded both in the relative abundance of core histones and variants and particularly in the post-translational modification of these proteins. We determined the presence of such variants and covalent modifications in seven tissue types of the anuran Xenopus laevis, including oocyte, egg, sperm, early embryo equivalent (pronuclei incubated in egg extract), S3 neurula cells, A6 kidney cells, and erythrocytes. We first developed a new robust method for isolating the stored, predeposition histones from oocytes and eggs via chromatography on heparin-Sepharose, whereas we isolated chromatinized histones via conventional acid extraction. We identified two previously unknown H1 isoforms (H1fx and H1B.Sp) present on sperm chromatin. We immunoblotted this global collection of histones with many specific post-translational modification antibodies, including antibodies against methylated histone H3 on Lys(4), Lys(9), Lys(27), Lys(79), Arg(2), Arg(17), and Arg(26); methylated histone H4 on Lys(20); methylated H2A and H4 on Arg(3); acetylated H4 on Lys(5), Lys(8), Lys(12), and Lys(16) and H3 on Lys(9) and Lys(14); and phosphorylated H3 on Ser(10) and H2A/H4 on Ser(1). Furthermore, we subjected a subset of these histones to two-dimensional gel analysis and subsequent immunoblotting and mass spectrometry to determine the global remodeling of histone modifications that occurs as development proceeds. Overall, our observations suggest that each metazoan cell type may have a unique histone modification signature correlated with its differentiation status. | | 18957438
 |
Phosphorylation of histone H2A by protein kinase C and identification of the phosphorylation site.
Takeuchi, F, et al.
J. Biochem., 111: 788-92 (1992)
1992
Zobrazit abstrakt
In regenerating rat liver, nuclear protein histone H2A was shown to be phosphorylated on its amino-terminal serine residue [Sung et al. (1971) J. Biol. Chem. 246, 1358-1364], but the protein kinase which phosphorylates this residue has not been identified. To evaluate the possibility that protein kinase C can phosphorylate this residue, calf thymus histone H2A was 32P-labeled by incubation with [gamma-32P]ATP and highly purified protein kinase C from rat brain in the presence of calcium and phospholipid. About 1 mol of 32P was incorporated per mol of histone H2A and the Km and apparent Vmax of the reaction were calculated to be 2.1 microM and 0.35 mumol/min/mg, respectively. So histone H2A seemed to be a good substrate for protein kinase C. Further, the proteolytic phosphopeptides of 32P-labeled histone H2A were isolated by means of a series of column chromatographies and analyzed for their amino acid compositions. Comparison of the data with the known primary structure of histone H2A revealed their amino acid sequence as 1Ser-Gly-Arg. These data suggest that protein kinase C may be a candidate for the protein kinase which phosphorylates the amino-terminal serine residue of histone H2A during the regeneration of rat liver. | | 1500420
 |