The PML-associated protein DEK regulates the balance of H3.3 loading on chromatin and is important for telomere integrity.
Ivanauskiene, K; Delbarre, E; McGhie, JD; Küntziger, T; Wong, LH; Collas, P
Genome research
24
1584-94
2014
Show Abstract
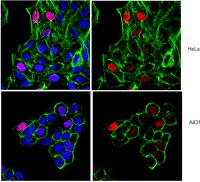
Histone variant H3.3 is deposited in chromatin at active sites, telomeres, and pericentric heterochromatin by distinct chaperones, but the mechanisms of regulation and coordination of chaperone-mediated H3.3 loading remain largely unknown. We show here that the chromatin-associated oncoprotein DEK regulates differential HIRA- and DAAX/ATRX-dependent distribution of H3.3 on chromosomes in somatic cells and embryonic stem cells. Live cell imaging studies show that nonnucleosomal H3.3 normally destined to PML nuclear bodies is re-routed to chromatin after depletion of DEK. This results in HIRA-dependent widespread chromatin deposition of H3.3 and H3.3 incorporation in the foci of heterochromatin in a process requiring the DAXX/ATRX complex. In embryonic stem cells, loss of DEK leads to displacement of PML bodies and ATRX from telomeres, redistribution of H3.3 from telomeres to chromosome arms and pericentric heterochromatin, induction of a fragile telomere phenotype, and telomere dysfunction. Our results indicate that DEK is required for proper loading of ATRX and H3.3 on telomeres and for telomeric chromatin architecture. We propose that DEK acts as a "gatekeeper" of chromatin, controlling chromatin integrity by restricting broad access to H3.3 by dedicated chaperones. Our results also suggest that telomere stability relies on mechanisms ensuring proper histone supply and routing. | 25049225
 |
HDA6 directly interacts with DNA methyltransferase MET1 and maintains transposable element silencing in Arabidopsis.
Liu, X; Yu, CW; Duan, J; Luo, M; Wang, K; Tian, G; Cui, Y; Wu, K
Plant physiology
158
119-29
2012
Show Abstract
The molecular mechanism of how the histone deacetylase HDA6 participates in maintaining transposable element (TE) silencing in Arabidopsis (Arabidopsis thaliana) is not yet defined. In this study, we show that a subset of TEs was transcriptionally reactivated and that TE reactivation was associated with elevated histone H3 and H4 acetylation as well as increased H3K4Me3 and H3K4Me2 in hda6 mutants. Decreased DNA methylation of the TEs was also detected in hda6 mutants, suggesting that HDA6 silences the TEs by regulating histone acetylation and methylation as well as the DNA methylation status of the TEs. Similarly, transcripts of some of these TEs were also increased in the methyltransferase1 (met1) mutant, with decreased DNA methylation. Furthermore, H4 acetylation, H3K4Me3, H3K4Me2, and H3K36Me2 were enriched at the coregulated TEs in the met1 and hda6 met1 mutants. Protein-protein interaction analysis indicated that HDA6 physically interacts with MET1 in vitro and in vivo, and further deletion analysis demonstrated that the carboxyl-terminal region of HDA6 and the bromo-adjacent homology domain of MET1 were responsible for the interaction. These results suggested that HDA6 and MET1 interact directly and act together to silence TEs by modulating DNA methylation, histone acetylation, and histone methylation status. Full Text Article | 21994348
 |
Differentially expressed genes are marked by histone 3 lysine 9 trimethylation in human cancer cells.
Wiencke, JK; Zheng, S; Morrison, Z; Yeh, RF
Oncogene
27
2412-21
2008
Show Abstract
Histone H3 lysine 9 trimethylation (H3K9Me3) has been associated with transcriptional repression, but recent findings implicate this chromatin modification in transcriptional activation and mRNA elongation by RNA polymerase II. Here, we applied immunoprecipitation (IP) with a custom DNA tiling microarray containing many transcription factors important in development and cancer (for example homeotic genes; N=683 total genes) to explore the relationship between H3K9Me3 and other histone modifications with the differential expression of genes. Cancer cell lines derived from different tissues (2 leukemia, 2 medulloblastoma) were characterized with IP antibodies to H3K9Me3, H3K4 dimethylation (H3K4Me2) and H3K9 acetylation (H3K9Ac). MV4-11 is known to overexpress the HOXA9 and MEIS1 genes, whereas D283 overexpresses the OTX2 homeobox gene. Gene expression was assessed by Affymetrix U133 array. Mapping the number and size of histone markings demonstrated significant colocalization of H3K9Ac and H3K4Me2 with H3K9Me3, indicating a pattern of putative 'activating' and 'repressive' markings. The median site size was 600-821 bp and 72-95% or 53-80% of chromatin signal sites were located within 1 kb or 500 bp of transcription start sites (TSS), respectively. A relatively small number of genes displayed additional H3K9Me3 sites in the 5'-region distant from the TSS. Comparing genes with modification sites to those without sites in their promoters confirmed the positive associations of H3K9Ac and H3K4Me2 with gene expression and revealed that H3K9Me3 is associated with active genes rather than being a repressive marking as previously thought. The positive regulatory effect of all three types of modifications were quantitatively correlated with site size, and applied to absolute gene expression within a single cell line as well as relative expression among pairs of cell lines. Extended patterns of H3K9Me3 upstream of some genes (for example HOXA9 and OTX2) may result from the action of multiple promoter elements. We found an inverse relationship between promoter DNA hypermethylation and H3K9Me3 in three studied genes (HOXA9, TMS1, RASSF1A). The localization of H3K9Me3 downstream of the TSSs of expressed genes and not within promoter regions of hypermethylated and silenced genes is consistent with the proposed coupling of H3K9Me3 with RNA polymerase II. Our results indicate a need for revising aspects of the histone code involving H3 lysine methylation. Awareness of H3K9Me3 as a mark of gene activity, not repression, is especially important for the classification of human cancer using chromatin and histone profiles. | 17968314
 |