Epigenetic regulation of traf2- and Nck-interacting kinase (TNIK) in polycystic ovary syndrome.
Li, D; Jiao, J; Zhou, YM; Wang, XX
American journal of translational research
7
1152-60
2015
Pokaż streszczenie
Emerging evidence has led to considerable interest in the role of Traf2- and Nck-interacting kinase (TNIK) in polycystic ovary syndrome (PCOS) development. However, the epigenetic mechanism regulating TNIK transcription remains largely unknown. Here, we show that (i) TNIK mRNA expression is significantly increased in PCOS ovarian tissues, compared to normal ovarian tissues; (ii) PCOS ovarian tissues exhibit a hypermethylation pattern at the cg10180092 site, (iii) and cg10180092 is the critical site for the transcriptional regulation of TNIK. Mechanistically, hypermethylated cg10180092 site-mediated loss of holocarboxylase synthetase (HLCS)-related H3K9me enrichment activated TNIK transcription in PCOS ovarian tissues. Notably, a substantial body of evidence indicates that DNA hypermethylation is an alternative mechanism for gene inactivation, and a new role for DNA hypermethylationmediated TNIK activating was observed in this study. This may improve our understanding of divergent transcriptional regulation in the initiation and progression of TNIK-related PCOS. | | | 26279758
 |
Quantification of histone H3 Lys27 trimethylation (H3K27me3) by high-throughput microscopy enables cellular large-scale screening for small-molecule EZH2 inhibitors.
Luense, S; Denner, P; Fernández-Montalván, A; Hartung, I; Husemann, M; Stresemann, C; Prechtl, S
Journal of biomolecular screening
20
190-201
2015
Pokaż streszczenie
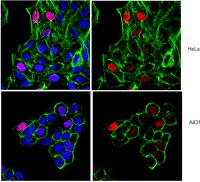
EZH2 inhibition can decrease global histone H3 lysine 27 trimethylation (H3K27me3) and thereby reactivates silenced tumor suppressor genes. Inhibition of EZH2 is regarded as an option for therapeutic cancer intervention. To identify novel small-molecule (SMOL) inhibitors of EZH2 in drug discovery, trustworthy cellular assays amenable for phenotypic high-throughput screening (HTS) are crucial. We describe a reliable approach that quantifies changes in global levels of histone modification marks using high-content analysis (HCA). The approach was validated in different cell lines by using small interfering RNA and SMOL inhibitors. By automation and miniaturization from a 384-well to 1536-well plate, we demonstrated its utility in conducting phenotypic HTS campaigns and assessing structure-activity relationships (SAR). This assay enables screening of SMOL EZH2 inhibitors and can advance the mechanistic understanding of H3K27me3 suppression, which is crucial with regard to epigenetic therapy. We observed that a decrease in global H3K27me3, induced by EZH2 inhibition, comprises two distinct mechanisms: (1) inhibition of de novo DNA methylation and (II) inhibition of dynamic, replication-independent H3K27me3 turnover. This report describes an HCA assay for primary HTS to identify, profile, and optimize cellular active SMOL inhibitors targeting histone methyltransferases, which could benefit epigenetic drug discovery. | | | 25409661
 |
Intracellular α-ketoglutarate maintains the pluripotency of embryonic stem cells.
Carey, BW; Finley, LW; Cross, JR; Allis, CD; Thompson, CB
Nature
518
413-6
2015
Pokaż streszczenie
The role of cellular metabolism in regulating cell proliferation and differentiation remains poorly understood. For example, most mammalian cells cannot proliferate without exogenous glutamine supplementation even though glutamine is a non-essential amino acid. Here we show that mouse embryonic stem (ES) cells grown under conditions that maintain naive pluripotency are capable of proliferation in the absence of exogenous glutamine. Despite this, ES cells consume high levels of exogenous glutamine when the metabolite is available. In comparison to more differentiated cells, naive ES cells utilize both glucose and glutamine catabolism to maintain a high level of intracellular α-ketoglutarate (αKG). Consequently, naive ES cells exhibit an elevated αKG to succinate ratio that promotes histone/DNA demethylation and maintains pluripotency. Direct manipulation of the intracellular αKG/succinate ratio is sufficient to regulate multiple chromatin modifications, including H3K27me3 and ten-eleven translocation (Tet)-dependent DNA demethylation, which contribute to the regulation of pluripotency-associated gene expression. In vitro, supplementation with cell-permeable αKG directly supports ES-cell self-renewal while cell-permeable succinate promotes differentiation. This work reveals that intracellular αKG/succinate levels can contribute to the maintenance of cellular identity and have a mechanistic role in the transcriptional and epigenetic state of stem cells. | | | 25487152
 |
Nascent chromatin capture proteomics determines chromatin dynamics during DNA replication and identifies unknown fork components.
Alabert, C; Bukowski-Wills, JC; Lee, SB; Kustatscher, G; Nakamura, K; de Lima Alves, F; Menard, P; Mejlvang, J; Rappsilber, J; Groth, A
Nature cell biology
16
281-93
2014
Pokaż streszczenie
To maintain genome function and stability, DNA sequence and its organization into chromatin must be duplicated during cell division. Understanding how entire chromosomes are copied remains a major challenge. Here, we use nascent chromatin capture (NCC) to profile chromatin proteome dynamics during replication in human cells. NCC relies on biotin-dUTP labelling of replicating DNA, affinity purification and quantitative proteomics. Comparing nascent chromatin with mature post-replicative chromatin, we provide association dynamics for 3,995 proteins. The replication machinery and 485 chromatin factors such as CAF-1, DNMT1 and SUV39h1 are enriched in nascent chromatin, whereas 170 factors including histone H1, DNMT3, MBD1-3 and PRC1 show delayed association. This correlates with H4K5K12diAc removal and H3K9me1 accumulation, whereas H3K27me3 and H3K9me3 remain unchanged. Finally, we combine NCC enrichment with experimentally derived chromatin probabilities to predict a function in nascent chromatin for 93 uncharacterized proteins, and identify FAM111A as a replication factor required for PCNA loading. Together, this provides an extensive resource to understand genome and epigenome maintenance. | | | 24561620
 |
Selective methylation of histone H3 variant H3.1 regulates heterochromatin replication.
Jacob, Y; Bergamin, E; Donoghue, MT; Mongeon, V; LeBlanc, C; Voigt, P; Underwood, CJ; Brunzelle, JS; Michaels, SD; Reinberg, D; Couture, JF; Martienssen, RA
Science (New York, N.Y.)
343
1249-53
2014
Pokaż streszczenie
Histone variants have been proposed to act as determinants for posttranslational modifications with widespread regulatory functions. We identify a histone-modifying enzyme that selectively methylates the replication-dependent histone H3 variant H3.1. The crystal structure of the SET domain of the histone H3 lysine-27 (H3K27) methyltransferase ARABIDOPSIS TRITHORAX-RELATED PROTEIN 5 (ATXR5) in complex with a H3.1 peptide shows that ATXR5 contains a bipartite catalytic domain that specifically "reads" alanine-31 of H3.1. Variation at position 31 between H3.1 and replication-independent H3.3 is conserved in plants and animals, and threonine-31 in H3.3 is responsible for inhibiting the activity of ATXR5 and its paralog, ATXR6. Our results suggest a simple model for the mitotic inheritance of the heterochromatic mark H3K27me1 and the protection of H3.3-enriched genes against heterochromatization during DNA replication. | | | 24626927
 |
Impact of human MLL/COMPASS and polycomb complexes on the DNA methylome.
Putiri, EL; Tiedemann, RL; Liu, C; Choi, JH; Robertson, KD
Oncotarget
5
6338-52
2014
Pokaż streszczenie
The correlation between DNA methylation and a subset of histone post-translational modifications (positive and negative) has hinted at an underlying regulatory crosstalk between histone marks and DNA methylation in patterning the human DNA methylome, an idea further supported by corresponding alterations to both histone marks and DNA methylation during malignant transformation. This study investigated the framework by which histone marks influence DNA methylation at a genome-wide level. Using RNAi in a pluripotent human embryonic carcinoma cell line we depleted essential components of the MLL/COMPASS, polycomb repressive complex 2 (PRC2), and PRC1 histone modifying complexes that establish, respectively, the post-translational modifications H3K4me3, H3K27me3, and H2AK119ub, and assayed the impact of the subsequent depletion of these marks on the DNA methylome. Absence of H2AK119ub resulted predominantly in hypomethylation across the genome. Depletion of H3K4me3 and, surprisingly, H3K27me3 caused CpG island hypermethylation at a subset of loci. Intriguingly, many promoters were co-regulated by all three histone marks, becoming hypermethylated with loss of H3K4me3 or H3K27me3 and hypomethylated with depletion of H2AK119ub, and many of these co-regulated loci were among those commonly targeted for aberrant hypermethylation in cancer. Taken together, our results elucidate novel roles for polycomb and MLL/COMPASS in regulating DNA methylation and define targets of this regulation. | Western Blotting | | 25071008
 |
Critical role of histone demethylase Jmjd3 in the regulation of CD4+ T-cell differentiation.
Li, Q; Zou, J; Wang, M; Ding, X; Chepelev, I; Zhou, X; Zhao, W; Wei, G; Cui, J; Zhao, K; Wang, HY; Wang, RF
Nature communications
5
5780
2014
Pokaż streszczenie
Epigenetic factors have been implicated in the regulation of CD4(+) T-cell differentiation. Jmjd3 plays a role in many biological processes, but its in vivo function in T-cell differentiation remains unknown. Here we report that Jmjd3 ablation promotes CD4(+) T-cell differentiation into Th2 and Th17 cells in the small intestine and colon, and inhibits T-cell differentiation into Th1 cells under different cytokine-polarizing conditions and in a Th1-dependent colitis model. Jmjd3 deficiency also restrains the plasticity of the conversion of Th2, Th17 or Treg cells to Th1 cells. The skewing of T-cell differentiation is concomitant with changes in the expression of key transcription factors and cytokines. H3K27me3 and H3K4me3 levels in Jmjd3-deficient cells are correlated with altered gene expression through interactions with specific transcription factors. Our results identify Jmjd3 as an epigenetic factor in T-cell differentiation via changes in histone methylation and target gene expression. | Western Blotting | | 25531312
 |
Stage-dependent and locus-specific role of histone demethylase Jumonji D3 (JMJD3) in the embryonic stages of lung development.
Li, Q; Wang, HY; Chepelev, I; Zhu, Q; Wei, G; Zhao, K; Wang, RF
PLoS genetics
10
e1004524
2014
Pokaż streszczenie
Histone demethylases have emerged as important players in developmental processes. Jumonji domain containing-3 (Jmjd3) has been identified as a key histone demethylase that plays a critical role in the regulation of gene expression; however, the in vivo function of Jmjd3 in embryonic development remains largely unknown. To this end, we generated Jmjd3 global and conditional knockout mice. Global deletion of Jmjd3 induces perinatal lethality associated with defective lung development. Tissue and stage-specific deletion revealed that Jmjd3 is dispensable in the later stage of embryonic lung development. Jmjd3 ablation downregulates the expression of genes critical for lung development and function, including AQP-5 and SP-B. Jmjd3-mediated alterations in gene expression are associated with locus-specific changes in the methylation status of H3K27 and H3K4. Furthermore, Jmjd3 is recruited to the SP-B promoter through interactions with the transcription factor Nkx2.1 and the epigenetic protein Brg1. Taken together, these findings demonstrate that Jmjd3 plays a stage-dependent and locus-specific role in the mouse lung development. Our study provides molecular insights into the mechanisms by which Jmjd3 regulates target gene expression in the embryonic stages of lung development. | | | 25079229
 |
Biphasic euchromatin-to-heterochromatin transition on the KSHV genome following de novo infection.
Toth, Z; Brulois, K; Lee, HR; Izumiya, Y; Tepper, C; Kung, HJ; Jung, JU
PLoS pathogens
9
e1003813
2013
Pokaż streszczenie
The establishment of latency is an essential step for the life-long persistent infection and pathogenesis of Kaposi's sarcoma-associated herpesvirus (KSHV). While the KSHV genome is chromatin-free in the virions, the viral DNA in latently infected cells has a chromatin structure with activating and repressive histone modifications that promote latent gene expression but suppress lytic gene expression. Here, we report a comprehensive epigenetic study of the recruitment of chromatin regulatory factors onto the KSHV genome during the pre-latency phase of KSHV infection. This demonstrates that the KSHV genome undergoes a biphasic chromatinization following de novo infection. Initially, a transcriptionally active chromatin (euchromatin), characterized by high levels of the H3K4me3 and acetylated H3K27 (H3K27ac) activating histone marks, was deposited on the viral episome and accompanied by the transient induction of a limited number of lytic genes. Interestingly, temporary expression of the RTA protein facilitated the increase of H3K4me3 and H3K27ac occupancy on the KSHV episome during de novo infection. Between 24-72 hours post-infection, as the levels of these activating histone marks declined on the KSHV genome, the levels of the repressive H3K27me3 and H2AK119ub histone marks increased concomitantly with the decline of lytic gene expression. Importantly, this transition to heterochromatin was dependent on both Polycomb Repressive Complex 1 and 2. In contrast, upon infection of human gingiva-derived epithelial cells, the KSHV genome underwent a transcription-active euchromatinization, resulting in efficient lytic gene expression. Our data demonstrate that the KSHV genome undergoes a temporally-ordered biphasic euchromatin-to-heterochromatin transition in endothelial cells, leading to latent infection, whereas KSHV preferentially adopts a transcriptionally active euchromatin in oral epithelial cells, resulting in lytic gene expression. Our results suggest that the differential epigenetic modification of the KSHV genome in distinct cell types is a potential determining factor for latent infection versus lytic replication of KSHV. | | | 24367262
 |
Epigenetic modifications and chromosome conformations of the beta globin locus throughout development.
Chang, KH; Fang, X; Wang, H; Huang, A; Cao, H; Yang, Y; Bonig, H; Stamatoyannopoulos, JA; Papayannopoulou, T
Stem cell reviews
9
397-407
2013
Pokaż streszczenie
Human embryonic stem cells provide an alternative to using human embryos for studying developmentally regulated gene expression. The co-expression of high levels of embryonic ε and fetal γ globin by the hESC-derived erythroblasts allows the interrogation of ε globin regulation at the transcriptional and epigenetic level which could only be attained previously by studying cell lines or transgenic mice. In this study, we compared the histone modifications across the β globin locus of the undifferentiated hESCs and hESC-, FL-, and mobilized PB CD34(+) cells-derived erythroblasts, which have distinct globin expression patterns corresponding to their developmental stages. We demonstrated that the histone codes employed by the β globin locus are conserved throughout development. Furthermore, in spite of the close proximity of the ε globin promoter, as compared to the β or γ globin promoter, with the LCR, a chromatin loop was also formed between the LCR and the active ε globin promoter, similar to the loop that forms between the β or γ globin promoters and the LCR, in contrary to the previously proposed tracking mechanism. | | | 22374078
 |