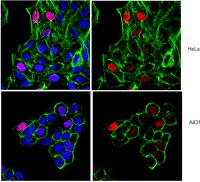
MAF1 represses CDKN1A through a Pol III-dependent mechanism.
Lee, YL; Li, YC; Su, CH; Chiao, CH; Lin, IH; Hsu, MT
eLife
4
e06283
2015
Afficher le résumé
MAF1 represses Pol III-mediated transcription by interfering with TFIIIB and Pol III. Herein, we found that MAF1 knockdown induced CDKN1A transcription and chromatin looping concurrently with Pol III recruitment. Simultaneous knockdown of MAF1 with Pol III or BRF1 (subunit of TFIIIB) diminished the activation and looping effect, which indicates that recruiting Pol III was required for activation of Pol II-mediated transcription and chromatin looping. Chromatin-immunoprecipitation analysis after MAF1 knockdown indicated enhanced binding of Pol III and BRF1, as well as of CFP1, p300, and PCAF, which are factors that mediate active histone marks, along with the binding of TATA binding protein (TBP) and POLR2E to the CDKN1A promoter. Simultaneous knockdown with Pol III abolished these regulatory events. Similar results were obtained for GDF15. Our results reveal a novel mechanism by which MAF1 and Pol III regulate the activity of a protein-coding gene transcribed by Pol II. | | | 26067234
 |
An anti-silencer- and SATB1-dependent chromatin hub regulates Rag1 and Rag2 gene expression during thymocyte development.
Hao, B; Naik, AK; Watanabe, A; Tanaka, H; Chen, L; Richards, HW; Kondo, M; Taniuchi, I; Kohwi, Y; Kohwi-Shigematsu, T; Krangel, MS
The Journal of experimental medicine
212
809-24
2015
Afficher le résumé
Rag1 and Rag2 gene expression in CD4(+)CD8(+) double-positive (DP) thymocytes depends on the activity of a distant anti-silencer element (ASE) that counteracts the activity of an intergenic silencer. However, the mechanistic basis for ASE activity is unknown. Here, we show that the ASE physically interacts with the distant Rag1 and Rag2 gene promoters in DP thymocytes, bringing the two promoters together to form an active chromatin hub. Moreover, we show that the ASE functions as a classical enhancer that can potently activate these promoters in the absence of the silencer or other locus elements. In thymocytes lacking the chromatin organizer SATB1, we identified a partial defect in Tcra gene rearrangement that was associated with reduced expression of Rag1 and Rag2 at the DP stage. SATB1 binds to the ASE and Rag promoters, facilitating inclusion of Rag2 in the chromatin hub and the loading of RNA polymerase II to both the Rag1 and Rag2 promoters. Our results provide a novel framework for understanding ASE function and demonstrate a novel role for SATB1 as a regulator of Rag locus organization and gene expression in DP thymocytes. | | | 25847946
 |
ZEB1-associated drug resistance in cancer cells is reversed by the class I HDAC inhibitor mocetinostat.
Meidhof, S; Brabletz, S; Lehmann, W; Preca, BT; Mock, K; Ruh, M; Schüler, J; Berthold, M; Weber, A; Burk, U; Lübbert, M; Puhr, M; Culig, Z; Wellner, U; Keck, T; Bronsert, P; Küsters, S; Hopt, UT; Stemmler, MP; Brabletz, T
EMBO molecular medicine
7
831-47
2015
Afficher le résumé
Therapy resistance is a major clinical problem in cancer medicine and crucial for disease relapse and progression. Therefore, the clinical need to overcome it, particularly for aggressive tumors such as pancreatic cancer, is very high. Aberrant activation of an epithelial-mesenchymal transition (EMT) and an associated cancer stem cell phenotype are considered a major cause of therapy resistance. Particularly, the EMT-activator ZEB1 was shown to confer stemness and resistance. We applied a systematic, stepwise strategy to interfere with ZEB1 function, aiming to overcome drug resistance. This led to the identification of both its target gene miR-203 as a major drug sensitizer and subsequently the class I HDAC inhibitor mocetinostat as epigenetic drug to interfere with ZEB1 function, restore miR-203 expression, repress stemness properties, and induce sensitivity against chemotherapy. Thereby, mocetinostat turned out to be more effective than other HDAC inhibitors, such as SAHA, indicating the relevance of the screening strategy. Our data encourage the application of mechanism-based combinations of selected epigenetic drugs with standard chemotherapy for the rational treatment of aggressive solid tumors, such as pancreatic cancer. | | | 25872941
 |
Hierarchical clustering of breast cancer methylomes revealed differentially methylated and expressed breast cancer genes.
Lin, IH; Chen, DT; Chang, YF; Lee, YL; Su, CH; Cheng, C; Tsai, YC; Ng, SC; Chen, HT; Lee, MC; Chen, HW; Suen, SH; Chen, YC; Liu, TT; Chang, CH; Hsu, MT
PloS one
10
e0118453
2015
Afficher le résumé
Oncogenic transformation of normal cells often involves epigenetic alterations, including histone modification and DNA methylation. We conducted whole-genome bisulfite sequencing to determine the DNA methylomes of normal breast, fibroadenoma, invasive ductal carcinomas and MCF7. The emergence, disappearance, expansion and contraction of kilobase-sized hypomethylated regions (HMRs) and the hypomethylation of the megabase-sized partially methylated domains (PMDs) are the major forms of methylation changes observed in breast tumor samples. Hierarchical clustering of HMR revealed tumor-specific hypermethylated clusters and differential methylated enhancers specific to normal or breast cancer cell lines. Joint analysis of gene expression and DNA methylation data of normal breast and breast cancer cells identified differentially methylated and expressed genes associated with breast and/or ovarian cancers in cancer-specific HMR clusters. Furthermore, aberrant patterns of X-chromosome inactivation (XCI) was found in breast cancer cell lines as well as breast tumor samples in the TCGA BRCA (breast invasive carcinoma) dataset. They were characterized with differentially hypermethylated XIST promoter, reduced expression of XIST, and over-expression of hypomethylated X-linked genes. High expressions of these genes were significantly associated with lower survival rates in breast cancer patients. Comprehensive analysis of the normal and breast tumor methylomes suggests selective targeting of DNA methylation changes during breast cancer progression. The weak causal relationship between DNA methylation and gene expression observed in this study is evident of more complex role of DNA methylation in the regulation of gene expression in human epigenetics that deserves further investigation. | | | 25706888
 |
Quantification of histone H3 Lys27 trimethylation (H3K27me3) by high-throughput microscopy enables cellular large-scale screening for small-molecule EZH2 inhibitors.
Luense, S; Denner, P; Fernández-Montalván, A; Hartung, I; Husemann, M; Stresemann, C; Prechtl, S
Journal of biomolecular screening
20
190-201
2015
Afficher le résumé
EZH2 inhibition can decrease global histone H3 lysine 27 trimethylation (H3K27me3) and thereby reactivates silenced tumor suppressor genes. Inhibition of EZH2 is regarded as an option for therapeutic cancer intervention. To identify novel small-molecule (SMOL) inhibitors of EZH2 in drug discovery, trustworthy cellular assays amenable for phenotypic high-throughput screening (HTS) are crucial. We describe a reliable approach that quantifies changes in global levels of histone modification marks using high-content analysis (HCA). The approach was validated in different cell lines by using small interfering RNA and SMOL inhibitors. By automation and miniaturization from a 384-well to 1536-well plate, we demonstrated its utility in conducting phenotypic HTS campaigns and assessing structure-activity relationships (SAR). This assay enables screening of SMOL EZH2 inhibitors and can advance the mechanistic understanding of H3K27me3 suppression, which is crucial with regard to epigenetic therapy. We observed that a decrease in global H3K27me3, induced by EZH2 inhibition, comprises two distinct mechanisms: (1) inhibition of de novo DNA methylation and (II) inhibition of dynamic, replication-independent H3K27me3 turnover. This report describes an HCA assay for primary HTS to identify, profile, and optimize cellular active SMOL inhibitors targeting histone methyltransferases, which could benefit epigenetic drug discovery. | | | 25409661
 |
T-cell receptor α enhancer is inactivated in αβ T lymphocytes.
del Blanco, B; Angulo, Ú; Krangel, MS; Hernández-Munain, C
Proceedings of the National Academy of Sciences of the United States of America
112
E1744-53
2015
Afficher le résumé
The Tcra enhancer (Eα) is essential for Tcra locus germ-line transcription and primary Vα-to-Jα recombination during thymocyte development. We found that Eα is inhibited late during thymocyte differentiation and in αβ T lymphocytes, indicating that it is not required to drive transcription of rearranged Tcra genes. Eα inactivation resulted in the disruption of functional long-range enhancer-promoter interactions and was associated with loss of Eα-dependent histone modifications at promoter and enhancer regions, and reduced expression and recruitment of E2A to the Eα enhanceosome in T cells. Enhancer activity could not be recovered by T-cell activation, by forced expression of E2A or by the up-regulation of this and other transcription factors in the context of T helper differentiation. Our results argue that the major function of Eα is to coordinate the formation of a chromatin hub that drives Vα and Jα germ-line transcription and primary rearrangements in thymocytes and imply the existence of an Eα-independent mechanism to activate transcription of the rearranged Tcra locus in αβ T cells. | Immunoprecipitation | | 25831496
 |
Distinct patterns of the histone marks associated with recruitment of the methionine chain-elongation pathway from leucine biosynthesis.
Xue, M; Long, J; Jiang, Q; Wang, M; Chen, S; Pang, Q; He, Y
Journal of experimental botany
66
805-12
2015
Afficher le résumé
Aliphatic glucosinolates (GLSs) are derived from chain-elongated methionine produced by an iterative three-step process, known to be evolutionarily recruited from leucine biosynthesis. The divergence of homologous genes between two pathways is mainly linked to the alterations in biochemical features. In this study, it was discovered that a distinct pattern of histone modifications is associated with and/or contributes to the divergence of the two pathways. In general, genes involved in leucine biosynthesis were robustly associated with H3k4me2 and H3K4me3. In contrast, despite the considerable abundances of H3K4me2 observed in some of genes involved in methionine chain elongation, H3K4me3 was completely missing. This H3K4m3-depleted pattern had no effect on gene transcription, whereas it seemingly co-evolved with the entire pathway of aliphatic GLS biosynthesis. The results reveal a novel association of the epigenetic marks with plant secondary metabolism, and may help to understand the recruitment of the methionine chain-elongation pathway from leucine biosynthesis. | | | 25428994
 |
ChIP-Seq analysis of the adult male mouse brain after developmental exposure to arsenic.
Tyler, CR; Weber, JA; Labrecque, M; Hessinger, JM; Edwards, JS; Allan, AM
Data in brief
5
248-54
2015
Afficher le résumé
Exposure to the common environmental contaminant arsenic impacts the epigenetic landscape, including DNA methylation and histone modifications, of several cell types. Developmental arsenic exposure (DAE) increases acetylation and methylation of histone proteins and the protein expression of several chromatin-modifying enzymes in the dentate gyrus (DG) subregion of the adult male mouse brain [26]. To complement and support these data, ChIP-Seq analysis of DNA associated with trimethylation of histone 3 lysine 4 (H3K4me3) derived from the adult male DG after DAE was performed. DAE induced differential H3K4me3 enrichment on genes in pathways associated with cellular development and growth, cell death and survival, and neurological disorders, particularly as they relate to cancer, in the adult male brain. Comparison of H3K4me3 enrichment in controls revealed mechanisms that are potentially lacking in arsenic-exposed animals, including neurotransmission, neuronal growth and development, hormonal regulation, protein synthesis, and cellular homeostasis. New pathways impacted by arsenic include cytoskeleton organization, cell signaling, and potential disruption of immune function and warrant further investigation using this DAE paradigm in the mouse brain. | | | 26543888
 |
The Epigenome of Schistosoma mansoni Provides Insight about How Cercariae Poise Transcription until Infection.
Roquis, D; Lepesant, JM; Picard, MA; Freitag, M; Parrinello, H; Groth, M; Emans, R; Cosseau, C; Grunau, C
PLoS neglected tropical diseases
9
e0003853
2015
Afficher le résumé
Chromatin structure can control gene expression and can define specific transcription states. For example, bivalent methylation of histone H3K4 and H3K27 is linked to poised transcription in vertebrate embryonic stem cells (ESC). It allows them to rapidly engage specific developmental pathways. We reasoned that non-vertebrate metazoans that encounter a similar developmental constraint (i.e. to quickly start development into a new phenotype) might use a similar system. Schistosomes are parasitic platyhelminthes that are characterized by passage through two hosts: a mollusk as intermediate host and humans or rodents as definitive host. During its development, the parasite undergoes drastic changes, most notable immediately after infection of the definitive host, i.e. during the transition from the free-swimming cercariae into adult worms.We used Chromatin Immunoprecipitation followed by massive parallel sequencing (ChIP-Seq) to analyze genome-wide chromatin structure of S. mansoni on the level of histone modifications (H3K4me3, H3K27me3, H3K9me3, and H3K9ac) in cercariae, schistosomula and adults (available at http://genome.univ-perp.fr). We saw striking differences in chromatin structure between the developmental stages, but most importantly we found that cercariae possess a specific combination of marks at the transcription start sites (TSS) that has similarities to a structure found in ESC. We demonstrate that in cercariae no transcription occurs, and we provide evidences that cercariae do not possess large numbers of canonical stem cells.We describe here a broad view on the epigenome of a metazoan parasite. Most notably, we find bivalent histone H3 methylation in cercariae. Methylation of H3K27 is removed during transformation into schistosomula (and stays absent in adults) and transcription is activated. In addition, shifts of H3K9 methylation and acetylation occur towards upstream and downstream of the transcriptional start site (TSS). We conclude that specific H3 modifications are a phylogenetically older and probably more general mechanism, i.e. not restricted to stem cells, to poise transcription. Since adult couples must form to cause the disease symptoms, changes in histone modifications appear to be crucial for pathogenesis and represent therefore a therapeutic target. | | | 26305466
 |
Geminivirus-encoded TrAP suppressor inhibits the histone methyltransferase SUVH4/KYP to counter host defense.
Castillo-González, C; Liu, X; Huang, C; Zhao, C; Ma, Z; Hu, T; Sun, F; Zhou, Y; Zhou, X; Wang, XJ; Zhang, X
eLife
4
e06671
2015
Afficher le résumé
Transcriptional gene silencing (TGS) can serve as an innate immunity against invading DNA viruses throughout Eukaryotes. Geminivirus code for TrAP protein to suppress the TGS pathway. Here, we identified an Arabidopsis H3K9me2 histone methyltransferase, Su(var)3-9 homolog 4/Kryptonite (SUVH4/KYP), as a bona fide cellular target of TrAP. TrAP interacts with the catalytic domain of KYP and inhibits its activity in vitro. TrAP elicits developmental anomalies phenocopying several TGS mutants, reduces the repressive H3K9me2 mark and CHH DNA methylation, and reactivates numerous endogenous KYP-repressed loci in vivo. Moreover, KYP binds to the viral chromatin and controls its methylation to combat virus infection. Notably, kyp mutants support systemic infection of TrAP-deficient Geminivirus. We conclude that TrAP attenuates the TGS of the viral chromatin by inhibiting KYP activity to evade host surveillance. These findings provide new insight on the molecular arms race between host antiviral defense and virus counter defense at an epigenetic level. | | | 26344546
 |