Dynamic loss of H2B ubiquitylation without corresponding changes in H3K4 trimethylation during myogenic differentiation.
Vethantham, V; Yang, Y; Bowman, C; Asp, P; Lee, JH; Skalnik, DG; Dynlacht, BD
Molecular and cellular biology
32
1044-55
2011
Abstract anzeigen
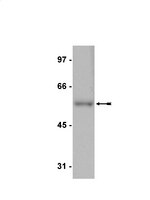
Ubiquitylation of H2B on lysine 120 (H2Bub) is associated with active transcriptional elongation. H2Bub has been implicated in histone cross talk and is generally regarded to be a prerequisite for trimethylation of histone 3 lysine 4 (H3K4me3) and H3K79 in both yeast and mammalian cells. We performed a genome-wide analysis of epigenetic marks during muscle differentiation, and strikingly, we observed a near-complete loss of H2Bub in the differentiated state. We examined the basis for global loss of this mark and found that the H2B ubiquitin E3 ligase, RNF20, was depleted from chromatin in differentiated myotubes, indicating that recruitment of this protein to genes substantially decreases upon differentiation. Remarkably, during the course of myogenic differentiation, we observed retention and acquisition of H3K4 trimethylation on a large number of genes in the absence of detectable H2Bub. The Set1 H3K4 trimethylase complex was efficiently recruited to a subset of genes in myotubes in the absence of detectable H2Bub, accounting in part for H3K4 trimethylation in myotubes. Our studies suggest that H3K4me3 deposition in the absence of detectable H2Bub in myotubes is mediated via Set1 and, perhaps, MLL complexes, whose recruitment does not require H2Bub. Thus, muscle cells represent a novel setting in which to explore mechanisms that regulate histone cross talk. | | 22252316
 |
CCN3/NOV small interfering RNA enhances fibrogenic gene expression in primary hepatic stellate cells and cirrhotic fat storing cell line CFSC.
Borkham-Kamphorst, E; van Roeyen, CR; Van de Leur, E; Floege, J; Weiskirchen, R
Journal of cell communication and signaling
6
11-25
2011
Abstract anzeigen
Nephroblastoma overexpressed gene encodes a matricellular protein (CCN3/NOV) of the CCN family, comprising CCN1 (CYR61), CCN2 (CTGF), CCN4 (WISP-1), CCN5 (WISP-2), and CCN6 (WISP-3). CCN proteins are involved in the regulation of mitosis, adhesion, apoptosis, extracellular matrix production, growth arrest and migration in multiple cell types. Compared to CCN2/CTGF, known as a profibrotic protein, the biological role of CCN3/NOV in liver fibrosis remains obscure. In this study we showed ccn3/nov mRNA to increase dramatically following hepatic stellate cell activation, reaching peak levels in fully transdifferentiated myofibroblasts. In models of experimental hepatic fibrosis, CCN3/NOV increased significantly at the mRNA and protein levels. CCN3/NOV was found mainly in non-parenchymal cells along the areas of tissue damage and repair. In the bile-duct ligation model, CCN3/NOV was localized mainly along portal tracts, while the repeated application of carbon tetrachloride resulted in CCN3/NOV expression mainly in the centrilobular areas. In contrast to CCN2/CTGF, the profibrotic cytokines platelet-derived growth factor-B and -D as well as transforming growth factor-β suppressed CCN3/NOV expression. In vitro, CCN3/NOV siRNA attenuated migration in the cirrhotic fat storing cell line CFSC well in line with in vivo findings that various types of cells expressing CCN3/NOV migrate into the area of tissue damage and regeneration. The suppression of CCN3/NOV enhanced expression of profibrotic marker proteins, such as α-smooth muscle actin, collagen type I, fibronectin, CCN2/CTGF and TIMP-1 in primary rat hepatic stellate cells and in CFSC. We further found that adenoviral overexpression of CCN2/CTGF suppressed CCN3/NOV expression, while the overexpression of CCN3/NOV as well as the suppression of CCN3/NOV by targeting siRNAs both resulted in enhanced CCN2/CTGF expression. These results indicate the complexity of CCN actions that are far beyond the classic Yin/Yang interplay. | | 21748432
 |
Functional interaction between E2F-4 and p130: evidence for distinct mechanisms underlying growth suppression by different retinoblastoma protein family members.
Vairo, G, et al.
Genes Dev., 9: 869-81 (1995)
1994
Abstract anzeigen
Little is known of the mechanisms controlling the G0/G1 transition of the cell cycle. The induction of immediate early gene expression, thought to be important for this process, suggests that the key factors controlling this transition preexist in quiescent cells. The E2F family of transcription factors likely play an important role in this process, because E2F DNA-binding activity exists in quiescent cells, and the induction of at least some immediate early genes requires intact E2F regulatory promoter sites. Here, we show that the major G0 E2F activity of primary human T cells, E2F-4, is stably bound to the p130 pocket protein in association with a DP heterodimerization partner. p130-E2F-4 binding has functional implications because p130 effectively suppressed E2F-4-mediated trans-activation, and coexpression of E2F4 overcame p130-mediated G1 arrest more efficiently than RB-induced G1 blockade. Conversely, E2F-1 overrode an RB-induced G1 block more efficiently than E2F-4. Thus, p130 and RB appear to induce cell cycle arrest via biochemically distinct mechanisms that involve different E2F family members. | | 7705662
 |
E2F-4, a new member of the E2F transcription factor family, interacts with p107.
Ginsberg, D, et al.
Genes Dev., 8: 2665-79 (1994)
1993
Abstract anzeigen
The E2F family of transcription factors has been implicated in the regulation of cell proliferation, and E2F-binding sites are present in the promoters of several growth-regulating genes. E2F family members are functionally regulated, in part, by complex formation with one or more members of the nuclear pocket protein family, RB, p107, and p130. Pocket protein regulation of E2F likely contributes to normal cellular growth control. While the three cloned species of E2F, E2F-1, E2F-2, and E2F-3, are known to be targets of RB interaction, no E2F species has yet been shown to be a specific p107 or p130 target. Here, we describe the cloning of a new member of the E2F family, E2F-4, which forms heterodimers with a member(s) of the DP family and, unlike some family members, is present throughout the cell cycle and appears to be a differentially phosphorylated p107-binding partner. p107 binding not only can be linked to the regulation of E2F-4 transcriptional activity, but also to suppression of the ability of E2F-4 to transform an immortalized rodent cell line. | | 7958924
 |