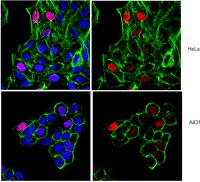
Naive pluripotency is associated with global DNA hypomethylation.
Leitch, HG; McEwen, KR; Turp, A; Encheva, V; Carroll, T; Grabole, N; Mansfield, W; Nashun, B; Knezovich, JG; Smith, A; Surani, MA; Hajkova, P
Nature structural & molecular biology
20
311-6
2013
Show Abstract
Naive pluripotent embryonic stem cells (ESCs) and embryonic germ cells (EGCs) are derived from the preimplantation epiblast and primordial germ cells (PGCs), respectively. We investigated whether differences exist between ESCs and EGCs, in view of their distinct developmental origins. PGCs are programmed to undergo global DNA demethylation; however, we find that EGCs and ESCs exhibit equivalent global DNA methylation levels. Inhibition of MEK and Gsk3b by 2i conditions leads to pronounced reduction in DNA methylation in both cell types. This is driven by Prdm14 and is associated with downregulation of Dnmt3a and Dnmt3b. However, genomic imprints are maintained in 2i, and we report derivation of EGCs with intact genomic imprints. Collectively, our findings establish that culture in 2i instills a naive pluripotent state with a distinctive epigenetic configuration that parallels molecular features observed in both the preimplantation epiblast and nascent PGCs. | Western Blotting | 23416945
 |
Ornithine decarboxylase antizyme induces hypomethylation of genome DNA and histone H3 lysine 9 dimethylation (H3K9me2) in human oral cancer cell line.
Yamamoto, D; Shima, K; Matsuo, K; Nishioka, T; Chen, CY; Hu, GF; Sasaki, A; Tsuji, T
PloS one
5
e12554
2010
Show Abstract
Methylation of CpG islands of genome DNA and lysine residues of histone H3 and H4 tails regulates gene transcription. Inhibition of polyamine synthesis by ornithine decarboxylase antizyme-1 (OAZ) in human oral cancer cell line resulted in accumulation of decarboxylated S-adenosylmethionine (dcSAM), which acts as a competitive inhibitor of methylation reactions. We anticipated that accumulation of dcSAM impaired methylation reactions and resulted in hypomethylation of genome DNA and histone tails.Global methylation state of genome DNA and lysine residues of histone H3 and H4 tails were assayed by Methylation by Isoschizomers (MIAMI) method and western blotting, respectively, in the presence or absence of OAZ expression. Ectopic expression of OAZ mediated hypomethylation of CpG islands of genome DNA and histone H3 lysine 9 dimethylation (H3K9me2). Protein level of DNA methyltransferase 3B (DNMT3B) and histone H3K9me specific methyltransferase G9a were down-regulated in OAZ transfectant.OAZ induced hypomethylation of CpG islands of global genome DNA and H3K9me2 by down-regulating DNMT3B and G9a protein level. Hypomethylation of CpG islands of genome DNA and histone H3K9me2 is a potent mechanism of induction of the genes related to tumor suppression and DNA double strand break repair. Full Text Article | Western Blotting | 20838441
 |
Mass spectrometry analysis of the variants of histone H3 and H4 of soybean and their post-translational modifications.
Wu, T; Yuan, T; Tsai, SN; Wang, C; Sun, SM; Lam, HM; Ngai, SM
BMC plant biology
9
98
2009
Show Abstract
Histone modifications and histone variants are of importance in many biological processes. To understand the biological functions of the global dynamics of histone modifications and histone variants in higher plants, we elucidated the variants and post-translational modifications of histones in soybean, a legume plant with a much bigger genome than that of Arabidopsis thaliana.In soybean leaves, mono-, di- and tri-methylation at Lysine 4, Lysine 27 and Lysine 36, and acetylation at Lysine 14, 18 and 23 were detected in HISTONE H3. Lysine 27 was prone to being mono-methylated, while tri-methylation was predominant at Lysine 36. We also observed that Lysine 27 methylation and Lysine 36 methylation usually excluded each other in HISTONE H3. Although methylation at HISTONE H3 Lysine 79 was not reported in A. thaliana, mono- and di-methylated HISTONE H3 Lysine 79 were detected in soybean. Besides, acetylation at Lysine 8 and 12 of HISTONE H4 in soybean were identified. Using a combination of mass spectrometry and nano-liquid chromatography, two variants of HISTONE H3 were detected and their modifications were determined. They were different at positions of A31F41S87S90 (HISTONE variant H3.1) and T31Y41H87L90 (HISTONE variant H3.2), respectively. The methylation patterns in these two HISTONE H3 variants also exhibited differences. Lysine 4 and Lysine 36 methylation were only detected in HISTONE H3.2, suggesting that HISTONE variant H3.2 might be associated with actively transcribing genes. In addition, two variants of histone H4 (H4.1 and H4.2) were also detected, which were missing in other organisms. In the histone variant H4.1 and H4.2, the amino acid 60 was isoleucine and valine, respectively.This work revealed several distinct variants of soybean histone and their modifications that were different from A. thaliana, thus providing important biological information toward further understanding of the histone modifications and their functional significance in higher plants. | Western Blotting | 19643030
 |
The Iws1:Spt6:CTD complex controls cotranscriptional mRNA biosynthesis and HYPB/Setd2-mediated histone H3K36 methylation.
Yoh, SM; Lucas, JS; Jones, KA
Genes & development
22
3422-34
2008
Show Abstract
Many steps in gene expression and mRNA biosynthesis are coupled to transcription elongation and organized through the C-terminal domain (CTD) of the large subunit of RNA polymerase II (RNAPII). We showed recently that Spt6, a transcription elongation factor and histone H3 chaperone, binds to the Ser2P CTD and recruits Iws1 and the REF1/Aly mRNA export adaptor to facilitate mRNA export. Here we show that Iws1 also recruits the HYPB/Setd2 histone methyltransferase to the RNAPII elongation complex and is required for H3K36 trimethylation (H3K36me3) across the transcribed region of the c-Myc, HIV-1, and PABPC1 genes in vivo. Interestingly, knockdown of either Iws1 or HYPB/Setd2 also enhanced H3K27me3 at the 5' end of the PABPC1 gene, and depletion of Iws1, but not HYPB/Setd2, increased histone acetylation across the coding regions at the HIV-1 and PABPC1 genes in vivo. Knockdown of HYPB/Setd2, like Iws1, induced bulk HeLa poly(A)+ mRNAs to accumulate in the nucleus. In vitro, recombinant Spt6 binds selectively to a stretch of uninterrupted consensus repeats located in the N-terminal half of the CTD and recruits Iws1. Thus Iws1 connects two distinct CTD-binding proteins, Spt6 and HYPB/Setd2, in a megacomplex that affects mRNA export as well as the histone modification state of active genes. | | 19141475
 |