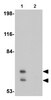
H3K9 methyltransferase G9a negatively regulates UHRF1 transcription during leukemia cell differentiation.
Kim, KB; Son, HJ; Choi, S; Hahm, JY; Jung, H; Baek, HJ; Kook, H; Hahn, Y; Kook, H; Seo, SB
Nucleic acids research
43
3509-23
2015
Mostrar resumen
Histone H3K9 methyltransferase (HMTase) G9a-mediated transcriptional repression is a major epigenetic silencing mechanism. UHRF1 (ubiquitin-like with PHD and ring finger domains 1) binds to hemimethylated DNA and plays an essential role in the maintenance of DNA methylation. Here, we provide evidence that UHRF1 is transcriptionally downregulated by H3K9 HMTase G9a. We found that increased expression of G9a along with transcription factor YY1 specifically represses UHRF1 transcription during TPA-mediated leukemia cell differentiation. Using ChIP analysis, we found that UHRF1 was among the transcriptionally silenced genes during leukemia cell differentiation. Using a DNA methylation profiling array, we discovered that the UHRF1 promoter was hypomethylated in samples from leukemia patients, further supporting its overexpression and oncogenic activity. Finally, we showed that G9a regulates UHRF1-mediated H3K23 ubiquitination and proper DNA replication maintenance. Therefore, we propose that H3K9 HMTase G9a is a specific epigenetic regulator of UHRF1. | | | 25765655
 |
Modulation of chromatin remodelling induced by the freshwater cyanotoxin cylindrospermopsin in human intestinal caco-2 cells.
Huguet, A; Hatton, A; Villot, R; Quenault, H; Blanchard, Y; Fessard, V
PloS one
9
e99121
2014
Mostrar resumen
Cylindrospermopsin (CYN) is a cyanotoxin that has been recognised as an emerging potential public health risk. Although CYN toxicity has been demonstrated, the mechanisms involved have not been fully characterised. To identify some key pathways related to this toxicity, we studied the transcriptomic profile of human intestinal Caco-2 cells exposed to a sub-toxic concentration of CYN (1.6 µM for 24hrs) using a non-targeted approach. CYN was shown to modulate different biological functions which were related to growth arrest (with down-regulation of cdkn1a and uhrf1 genes), and DNA recombination and repair (with up-regulation of aptx and pms2 genes). Our main results reported an increased expression of some histone-modifying enzymes (histone acetyl and methyltransferases MYST1, KAT5 and EHMT2) involved in chromatin remodelling, which is essential for initiating transcription. We also detected greater levels of acetylated histone H2A (Lys5) and dimethylated histone H3 (Lys4), two products of these enzymes. In conclusion, CYN overexpressed proteins involved in DNA damage repair and transcription, including modifications of nucleosomal histones. Our results highlighted some new cell processes induced by CYN. | | | 24921660
 |
Identification by high-throughput imaging of the histone methyltransferase EHMT2 as an epigenetic regulator of VEGFA alternative splicing.
Salton, M; Voss, TC; Misteli, T
Nucleic acids research
42
13662-73
2014
Mostrar resumen
Recent evidence points to a role of chromatin in regulation of alternative pre-mRNA splicing (AS). In order to identify novel chromatin regulators of AS, we screened an RNAi library of chromatin proteins using a cell-based high-throughput in vivo assay. We identified a set of chromatin proteins that regulate AS. Using simultaneous genome-wide expression and AS analysis, we demonstrate distinct and non-overlapping functions of these chromatin modifiers on transcription and AS. Detailed mechanistic characterization of one dual function chromatin modifier, the H3K9 methyltransferase EHMT2 (G9a), identified VEGFA as a major chromatin-mediated AS target. Silencing of EHMT2, or its heterodimer partner EHMT1, affects AS by promoting exclusion of VEGFA exon 6a, but does not alter total VEGFA mRNA levels. The epigenetic regulatory mechanism of AS by EHMT2 involves an adaptor system consisting of the chromatin modulator HP1γ, which binds methylated H3K9 and recruits splicing regulator SRSF1. The epigenetic regulation of VEGFA is physiologically relevant since EHMT2 is transcriptionally induced in response to hypoxia and triggers concomitant changes in AS of VEGFA. These results characterize a novel epigenetic regulatory mechanism of AS and they demonstrate separate roles of epigenetic modifiers in transcription and alternative splicing. | | | 25414343
 |
A chromatin activity-based chemoproteomic approach reveals a transcriptional repressome for gene-specific silencing.
Liu, C; Yu, Y; Liu, F; Wei, X; Wrobel, JA; Gunawardena, HP; Zhou, L; Jin, J; Chen, X
Nature communications
5
5733
2014
Mostrar resumen
Immune cells develop endotoxin tolerance (ET) after prolonged stimulation. ET increases the level of a repression mark H3K9me2 in the transcriptionally silent chromatin specifically associated with pro-inflammatory genes. However, it is not clear what proteins are functionally involved in this process. Here we show that a novel chromatin activity-based chemoproteomic (ChaC) approach can dissect the functional chromatin protein complexes that regulate ET-associated inflammation. Using UNC0638 that binds the enzymatically active H3K9-specific methyltransferase G9a/GLP, ChaC reveals that G9a is constitutively active at a G9a-dependent mega-dalton repressome in primary endotoxin-tolerant macrophages. G9a/GLP broadly impacts the ET-specific reprogramming of the histone code landscape, chromatin remodelling and the activities of select transcription factors. We discover that the G9a-dependent epigenetic environment promotes the transcriptional repression activity of c-Myc for gene-specific co-regulation of chronic inflammation. ChaC may also be applicable to dissect other functional protein complexes in the context of phenotypic chromatin architectures. | | | 25502336
 |
Profiling genome-wide chromatin methylation with engineered posttranslation apparatus within living cells.
Wang, R; Islam, K; Liu, Y; Zheng, W; Tang, H; Lailler, N; Blum, G; Deng, H; Luo, M
Journal of the American Chemical Society
135
1048-56
2013
Mostrar resumen
Protein methyltransferases (PMTs) have emerged as important epigenetic regulators in myriad biological processes in both normal physiology and disease conditions. However, elucidating PMT-regulated epigenetic processes has been hampered by ambiguous knowledge about in vivo activities of individual PMTs particularly because of their overlapping but nonredundant functions. To address limitations of conventional approaches in mapping chromatin modification of specific PMTs, we have engineered the chromatin-modifying apparatus and formulated a novel technology, termed clickable chromatin enrichment with parallel DNA sequencing (CliEn-seq), to probe genome-wide chromatin modification within living cells. The three-step approach of CliEn-seq involves in vivo synthesis of S-adenosyl-L-methionine (SAM) analogues from cell-permeable methionine analogues by engineered SAM synthetase (methionine adenosyltransferase or MAT), in situ chromatin modification by engineered PMTs, subsequent enrichment and sequencing of the uniquely modified chromatins. Given critical roles of the chromatin-modifying enzymes in epigenetics and structural similarity among many PMTs, we envision that the CliEn-seq technology is generally applicable in deciphering chromatin methylation events of individual PMTs in diverse biological settings. | Immunoprecipitation | | 23244065
 |
Gossypol and an HMT G9a inhibitor act in synergy to induce cell death in pancreatic cancer cells.
Yuan, Y; Tang, AJ; Castoreno, AB; Kuo, SY; Wang, Q; Kuballa, P; Xavier, R; Shamji, AF; Schreiber, SL; Wagner, BK
Cell death & disease
4
e690
2013
Mostrar resumen
The histone methyltransferase G9a is overexpressed in a variety of cancer types, including pancreatic adenocarcinoma, and promotes tumor invasiveness and metastasis. We recently reported the discovery of BRD4770, a small-molecule inhibitor of G9a that induces senescence in PANC-1 cells. We observed that the cytotoxic effects of BRD4770 were dependent on genetic background, with cell lines lacking functional p53 being relatively resistant to compound treatment. To understand the mechanism of genetic selectivity, we used two complementary screening approaches to identify enhancers of BRD4770. The natural product and putative BH3 mimetic gossypol enhanced the cytotoxicity of BRD4770 in a synergistic manner in p53-mutant PANC-1 cells but not in immortalized non-tumorigenic pancreatic cells. The combination of gossypol and BRD4770 increased LC3-II levels and the autophagosome number in PANC-1 cells, and the compound combination appears to act in a BNIP3 (B-cell lymphoma 2 19-kDa interacting protein)-dependent manner, suggesting that these compounds act together to induce autophagy-related cell death in pancreatic cancer cells. | | | 23807219
 |
REST-dependent epigenetic remodeling promotes the developmental switch in synaptic NMDA receptors.
Rodenas-Ruano, A; Chávez, AE; Cossio, MJ; Castillo, PE; Zukin, RS
Nature neuroscience
15
1382-90
2011
Mostrar resumen
NMDA receptors (NMDARs) are critical to synaptogenesis, neural circuitry and higher cognitive functions. A hallmark feature of NMDARs is an early postnatal developmental switch from those containing primarily GluN2B to primarily GluN2A subunits. Although the switch in phenotype has been an area of intense interest for two decades, the mechanisms that trigger it and the link between experience and the switch are unclear. Here we show a new role for the transcriptional repressor REST in the developmental switch of synaptic NMDARs. REST is activated at a critical window of time and acts via epigenetic remodeling to repress Grin2b expression and alter NMDAR properties at rat hippocampal synapses. Knockdown of REST in vivo prevented the decline in GluN2B and developmental switch in NMDARs. Maternal deprivation impaired REST activation and acquisition of the mature NMDAR phenotype. Thus, REST is essential for experience-dependent fine-tuning of genes involved in synaptic plasticity. | | | 22960932
 |
Negative regulation of JAK2 by H3K9 methyltransferase G9a in leukemia.
Son, HJ; Kim, JY; Hahn, Y; Seo, SB
Molecular and cellular biology
32
3681-94
2011
Mostrar resumen
Histone methylation at specific lysine residues is a crucial regulatory process in transcriptional regulation. Using chromatin immunoprecipitation with microarray technology (ChIP-chip) analysis, we found that the H3K9-me2 target gene JAK2 was an important factor during differentiation of the HL-60 promyelocytic leukemia cell line by all-trans-retinoic acid (ATRA) treatment. Here, we report that the H3K9 methyltransferase G9a negatively regulated JAK2 transcription in histone methyltransferase activity and in a YY1-dependent manner during ATRA-mediated leukemia cell differentiation. We found that G9a knockdown repressed ATRA-mediated HL-60 cell differentiation. We demonstrated that G9a interacts with YY1 and is recruited to the JAK2 promoter along with corepressors, including histone deacetylase, that induced H3K9-me2. Repression of JAK2 transcription by G9a decreased H3Y41 phosphorylation and promoted inhibition of the recently identified JAK2-H3Y41P-HP1α pathway-mediated leukemogenesis. | Fluorescence Activated Cell Sorting (FACS) | | 22801367
 |
Induction of the RNA regulator LIN28A is required for the growth and pathogenesis of RESTless breast tumors.
Gunsalus, KT; Wagoner, MP; Meyer, K; Potter, WB; Schoenike, B; Kim, S; Alexander, CM; Friedl, A; Roopra, A
Cancer research
72
3207-16
2011
Mostrar resumen
The transcription factor RE1 silencing transcription factor (REST) is lost in approximately 20% of breast cancers. Although it is known that these RESTless tumors are highly aggressive and include all tumor subtypes, the underlying tumorigenic mechanisms remain unknown. In this study, we show that loss of REST results in upregulation of LIN28A, a known promoter of tumor development, in breast cancer cell lines and human breast tumors. We found that LIN28A was a direct transcriptional target of REST in cancer cells and that loss of REST resulted in increased LIN28A expression and enhanced tumor growth both in vitro and in vivo, effects that were dependent on heightened LIN28A expression. Tumors lacking REST expression were locally invasive, consistent with the increased lymph node involvement observed in human RESTless tumors. Clinically, human RESTless breast tumors also displayed significantly enhanced LIN28A expression when compared with non-RESTless tumors. Our findings therefore show a critical role for the REST-LIN28A axis in tumor aggression and suggest a causative relationship between REST loss and tumorigenicity in vivo. | Western Blotting | | 22532168
 |
Dnmt3a1 upregulates transcription of distinct genes and targets chromosomal gene clusters for epigenetic silencing in mouse embryonic stem cells.
Kotini, AG; Mpakali, A; Agalioti, T
Molecular and cellular biology
31
1577-92
2010
Mostrar resumen
Dnmt3a1 and Dnmt3a2 are two de novo DNA methyltransferases expressed in mouse embryonic stem cells (mESCs). They differ in that a 219-amino-acid (aa) amino (N)-terminal noncatalytic domain is present only in Dnmt3a1. Here, we examined the unique functions of Dnmt3a1 in mESCs by targeting the coding sequence of the Dnmt3a1 N-terminal domain tagged with enhanced green fluorescent protein (GFP) for insertion into the mouse Rosa26 locus. Using these targeted cells (GFP-3a1Nter), we showed that Dnmt3a1 was efficiently recruited to the silenced Oct3/4 and activated Vtn (vitronectin) gene promoters via its unique N-terminal domain. This recruitment affected the two genes in contrasting ways, compromising Oct3/4 gene promoter DNA methylation to prevent consolidation of the silent state while significantly reducing Vtn transcription. We used this negative effect of the Dnmt3a1 N-terminal domain to investigate the extent of transcriptional regulation by Dnmt3a1 in mESCs by using microarrays. A small group of all-trans retinoic acid (tRA)-inducible genes had lower transcript levels in GFP-3a1Nter cells than in wild-type mESCs. Intriguingly, this group included genes that are important for fetal nutrition, placenta development, and metabolic functions and is enriched for a distinct set of imprinted genes. We also identified a larger group of genes that showed higher transcript levels in the GFP-3a1Nter-expressing cells than in wild-type mESCs, including pluripotency factors and key regulators of primordial germ cell differentiation. Thus, Dnmt3a1 in mESCs functions primarily as a negative and to a lesser extent as a positive regulator of transcription. Our findings suggest that Dnmt3a1 positively affects transcription of specific genes at the promoter level and targets chromosomal domains to epigenetically silence gene clusters in mESCs. | | | 21262766
 |