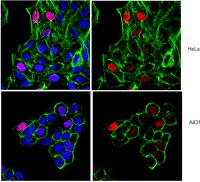
Chd1 is essential for the high transcriptional output and rapid growth of the mouse epiblast.
Guzman-Ayala, M; Sachs, M; Koh, FM; Onodera, C; Bulut-Karslioglu, A; Lin, CJ; Wong, P; Nitta, R; Song, JS; Ramalho-Santos, M
Development (Cambridge, England)
142
118-27
2015
显示摘要
The pluripotent mammalian epiblast undergoes unusually fast cell proliferation. This rapid growth is expected to generate a high transcriptional demand, but the underlying mechanisms remain unknown. We show here that the chromatin remodeler Chd1 is required for transcriptional output and development of the mouse epiblast. Chd1(-/-) embryos exhibit proliferation defects and increased apoptosis, are smaller than controls by E5.5 and fail to grow, to become patterned or to gastrulate. Removal of p53 allows progression of Chd1(-/-) mutants only to E7.0-8.0, highlighting the crucial requirement for Chd1 during early post-implantation development. Chd1(-/-) embryonic stem cells (ESCs) have a self-renewal defect and a genome-wide reduction in transcriptional output at both known mRNAs and intergenic transcripts. These transcriptional defects were only uncovered when cell number-normalized approaches were used, and correlate with a lower engagement of RNAP II with transcribed genes in Chd1(-/-) ESCs. We further show that Chd1 directly binds to ribosomal DNA, and that both Chd1(-/-) epiblast cells in vivo and ESCs in vitro express significantly lower levels of ribosomal RNA. In agreement with these findings, mutant cells in vivo and in vitro exhibit smaller and more elongated nucleoli. Thus, the RNA output by both Pol I and II is reduced in Chd1(-/-) cells. Our data indicate that Chd1 promotes a globally elevated transcriptional output required to sustain the distinctly rapid growth of the mouse epiblast. | | | 25480920
 |
Dopamine signaling leads to loss of Polycomb repression and aberrant gene activation in experimental parkinsonism.
Södersten, E; Feyder, M; Lerdrup, M; Gomes, AL; Kryh, H; Spigolon, G; Caboche, J; Fisone, G; Hansen, K
PLoS genetics
10
e1004574
2014
显示摘要
Polycomb group (PcG) proteins bind to and repress genes in embryonic stem cells through lineage commitment to the terminal differentiated state. PcG repressed genes are commonly characterized by the presence of the epigenetic histone mark H3K27me3, catalyzed by the Polycomb repressive complex 2. Here, we present in vivo evidence for a previously unrecognized plasticity of PcG-repressed genes in terminally differentiated brain neurons of parkisonian mice. We show that acute administration of the dopamine precursor, L-DOPA, induces a remarkable increase in H3K27me3S28 phosphorylation. The induction of the H3K27me3S28p histone mark specifically occurs in medium spiny neurons expressing dopamine D1 receptors and is dependent on Msk1 kinase activity and DARPP-32-mediated inhibition of protein phosphatase-1. Chromatin immunoprecipitation (ChIP) experiments showed that increased H3K27me3S28p was accompanied by reduced PcG binding to regulatory regions of genes. An analysis of the genome wide distribution of L-DOPA-induced H3K27me3S28 phosphorylation by ChIP sequencing (ChIP-seq) in combination with expression analysis by RNA-sequencing (RNA-seq) showed that the induction of H3K27me3S28p correlated with increased expression of a subset of PcG repressed genes. We found that induction of H3K27me3S28p persisted during chronic L-DOPA administration to parkisonian mice and correlated with aberrant gene expression. We propose that dopaminergic transmission can activate PcG repressed genes in the adult brain and thereby contribute to long-term maladaptive responses including the motor complications, or dyskinesia, caused by prolonged administration of L-DOPA in Parkinson's disease. | Western Blotting | Mouse | 25254549
 |
Alternative meiotic chromatid segregation in the holocentric plant Luzula elegans.
Heckmann, S; Jankowska, M; Schubert, V; Kumke, K; Ma, W; Houben, A
Nature communications
5
4979
2014
显示摘要
Holocentric chromosomes occur in a number of independent eukaryotic lineages. They form holokinetic kinetochores along the entire poleward chromatid surfaces, and owing to this alternative chromosome structure, species with holocentric chromosomes cannot use the two-step loss of cohesion during meiosis typical for monocentric chromosomes. Here we show that the plant Luzula elegans maintains a holocentric chromosome architecture and behaviour throughout meiosis, and in contrast to monopolar sister centromere orientation, the unfused holokinetic sister centromeres behave as two distinct functional units during meiosis I, resulting in sister chromatid separation. Homologous non-sister chromatids remain terminally linked after metaphase I, by satellite DNA-enriched chromatin threads, until metaphase II. They then separate at anaphase II. Thus, an inverted sequence of meiotic sister chromatid segregation occurs. This alternative meiotic process is most likely one possible adaptation to handle a holocentric chromosome architecture and behaviour during meiosis. | | | 25296379
 |
Genome-wide phosphoacetylation of histone H3 at Drosophila enhancers and promoters.
Kellner, WA; Ramos, E; Van Bortle, K; Takenaka, N; Corces, VG
Genome research
22
1081-8
2012
显示摘要
Transcription regulation is mediated by enhancers that bind sequence-specific transcription factors, which in turn interact with the promoters of the genes they control. Here, we show that the JIL-1 kinase is present at both enhancers and promoters of ecdysone-induced Drosophila genes, where it phosphorylates the Ser10 and Ser28 residues of histone H3. JIL-1 is also required for CREB binding protein (CBP)-mediated acetylation of Lys27, a well-characterized mark of active enhancers. The presence of these proteins at enhancers and promoters of ecdysone-induced genes results in the establishment of the H3K9acS10ph and H3K27acS28ph marks at both regulatory sequences. These modifications are necessary for the recruitment of 14-3-3, a scaffolding protein capable of facilitating interactions between two simultaneously bound proteins. Chromatin conformation capture assays indicate that interaction between the enhancer and the promoter is dependent on the presence of JIL-1, 14-3-3, and CBP. Genome-wide analyses extend these conclusions to most Drosophila genes, showing that the presence of JIL-1, H3K9acS10ph, and H3K27acS28ph is a general feature of enhancers and promoters in this organism. | | | 22508764
 |
Histone H3 phosphorylation (Ser10, Ser28) and phosphoacetylation (K9S10) are differentially associated with gene expression in liver of rats treated in vivo with acute ethanol.
James, TT; Aroor, AR; Lim, RW; Shukla, SD
J Pharmacol Exp Ther
340
237-47
2012
显示摘要
The epigenetic histone modification by ethanol is emerging as one of the mechanisms for its deleterious effects in the liver. In this context, we have investigated the role of histone H3 phosphorylation at Ser10 (P-H3-Ser10), and Ser28 (P-H3-Ser28) in liver after acute ethanol treatment in vivo. Ethanol was administered intraperitoneally in male Sprague-Dawley rats. Ethanol dose-response (1-5 g/kg body weight) and time-course (1-4 h) experiments were conducted, and various parameters were monitored. Steatosis and necrosis (serum alanine aminotransferase) of the liver increased in 4 h, suggesting liver injury. There were differences between P-H3-Ser10 and P-H3-Ser28 at 1 h, with the latter being more sensitive to lower ethanol doses. It was noteworthy that phosphorylation of both serines disappeared at the highest dose used (5 g/kg). We also examined phosphoacetylation of histone H3 at K9S10 and observed a dramatic increase. The changes in histone H3 phosphorylation and phosphoacetylation were also accompanied with expression of early response genes (c-fos, c-jun, mitogen-activated protein kinase phosphatase-1). Chromatin immunoprecipitation assays in samples from 1.5 and 4 h of ethanol administration indicated that increased histone H3 phosphorylation at Ser28 was associated with the promoters of c-jun and plasminogen activator inhibitor-1. In conclusion, this study demonstrates for the first time that in vivo exposure of liver to acute ethanol induced phosphorylation and phosphoacetylation of histone H3, and these modifications are differentially involved in the mRNA expression of genes. | | | 22025646
 |
Epigenetic changes in the brain: measuring global histone modifications.
Rumbaugh G, Miller CA
Methods Mol Biol
670
263-74.
2011
显示摘要
Recent years have witnessed an explosion of research on the role of epigenetic modifications, such as DNA methylation and histone protein acetylation and phosphorylation, in neuroscience. These changes exert control over gene expression and have been shown to play important roles in a variety of neural processes, including learning and memory. We and others have also recently shown that epigenetic changes may contribute to neurodegenerative disorders, such as Alzheimer's disease. Western blot analysis with antibodies raised against specific histone modifications is a relatively simple technique able to reveal the type, location, and degree of histone posttranslational modifications produced by an experimental manipulation. Here we provide a step-by-step protocol for isolating histone proteins from tissue and measuring these posttranslational modifications. | | | 20967596
 |
Ptf1a/Rbpj complex inhibits ganglion cell fate and drives the specification of all horizontal cell subtypes in the chick retina.
E C Lelièvre,M Lek,H Boije,L Houille-Vernes,V Brajeul,A Slembrouck,J E Roger,J A Sahel,J M Matter,F Sennlaub,F Hallböök,O Goureau,X Guillonneau
Developmental biology
358
2011
显示摘要
During development, progenitor cells of the retina give rise to six principal classes of neurons and the Müller glial cells found within the adult retina. The pancreas transcription factor 1 subunit a (Ptf1a) encodes a basic-helix-loop-helix transcription factor necessary for the specification of horizontal cells and the majority of amacrine cell subtypes in the mouse retina. The Ptf1a-regulated genes and the regulation of Ptf1a activity by transcription cofactors during retinogenesis have been poorly investigated. Using a retrovirus-mediated gene transfer approach, we reported that Ptf1a was sufficient to promote the fates of amacrine and horizontal cells from retinal progenitors and inhibit retinal ganglion cell and photoreceptor differentiation in the chick retina. Both GABAergic H1 and non-GABAergic H3 horizontal cells were induced following the forced expression of Ptf1a. We describe Ptf1a as a strong, negative regulator of Atoh7 expression. Furthermore, the Rbpj-interacting domains of Ptf1a protein were required for its effects on cell fate specification. Together, these data provide a novel insight into the molecular basis of Ptf1a activity on early cell specification in the chick retina. | | | 21839069
 |
Coe genes are expressed in differentiating neurons in the central nervous system of protostomes.
Demilly, A; Simionato, E; Ohayon, D; Kerner, P; Garcès, A; Vervoort, M
PloS one
6
e21213
2011
显示摘要
Genes of the coe (collier/olfactory/early B-cell factor) family encode Helix-Loop-Helix transcription factors that are widely conserved in metazoans and involved in many developmental processes, neurogenesis in particular. Whereas their functions during vertebrate neural tube formation have been well documented, very little is known about their expression and role during central nervous system (CNS) development in protostomes. Here we characterized the CNS expression of coe genes in the insect Drosophila melanogaster and the polychaete annelid Platynereis dumerilii, which belong to different subgroups of protostomes and show strikingly different modes of development. In the Drosophila ventral nerve cord, we found that the Collier-expressing cells form a subpopulation of interneurons with diverse molecular identities and neurotransmitter phenotypes. We also demonstrate that collier is required for the proper differentiation of some interneurons belonging to the Eve-Lateral cluster. In Platynereis dumerilii, we cloned a single coe gene, Pdu-coe, and found that it is exclusively expressed in post mitotic neural cells. Using an original technique of in silico 3D registration, we show that Pdu-coe is co-expressed with many different neuronal markers and therefore that, like in Drosophila, its expression defines a heterogeneous population of neurons with diverse molecular identities. Our detailed characterization and comparison of coe gene expression in the CNS of two distantly-related protostomes suggest conserved roles of coe genes in neuronal differentiation in this clade. As similar roles have also been observed in vertebrates, this function was probably already established in the last common ancestor of all bilaterians. | | | 21695052
 |
Calpain-1 cleaves Rad21 to promote sister chromatid separation.
Panigrahi, AK; Zhang, N; Mao, Q; Pati, D
Molecular and cellular biology
31
4335-47
2011
显示摘要
Defining the mechanisms of chromosomal cohesion and dissolution of the cohesin complex from chromatids is important for understanding the chromosomal missegregation seen in many tumor cells. Here we report the identification of a novel cohesin-resolving protease and describe its role in chromosomal segregation. Sister chromatids are held together by cohesin, a multiprotein ring-like complex comprised of Rad21, Smc1, Smc3, and SA2 (or SA1). Cohesin is known to be removed from vertebrate chromosomes by two distinct mechanisms, namely, the prophase and anaphase pathways. First, PLK1-mediated phosphorylation of SA2 in prophase leads to release of cohesin from chromosome arms, leaving behind centromeric cohesins that continue to hold the sisters together. Then, at the onset of anaphase, activated separase cleaves the centromeric cohesin Rad21, thereby opening the cohesin ring and allowing the sister chromatids to separate. We report here that the calcium-dependent cysteine endopeptidase calpain-1 is a Rad21 peptidase and normally localizes to the interphase nuclei and chromatin. Calpain-1 cleaves Rad21 at L192, in a calcium-dependent manner. We further show that Rad21 cleavage by calpain-1 promotes separation of chromosome arms, which coincides with a calcium-induced partial loss of cohesin at several chromosomal loci. Engineered cleavage of Rad21 at the calpain-cleavable site without activation of calpain-1 can lead to a loss of sister chromatid cohesion. Collectively, our work reveals a novel function of calpain-1 and describes an additional pathway for sister chromatid separation in humans. | | | 21876002
 |
PcG complexes set the stage for epigenetic inheritance of gene silencing in early S phase before replication.
Lanzuolo, C; Lo Sardo, F; Diamantini, A; Orlando, V
PLoS genetics
7
e1002370
2011
显示摘要
Polycomb group (PcG) proteins are part of a conserved cell memory system that conveys epigenetic inheritance of silenced transcriptional states through cell division. Despite the considerable amount of information about PcG mechanisms controlling gene silencing, how PcG proteins maintain repressive chromatin during epigenome duplication is still unclear. Here we identified a specific time window, the early S phase, in which PcG proteins are recruited at BX-C PRE target sites in concomitance with H3K27me3 repressive mark deposition. Notably, these events precede and are uncoupled from PRE replication timing, which occurs in late S phase when most epigenetic signatures are reduced. These findings shed light on one of the key mechanisms for PcG-mediated epigenetic inheritance during S phase, suggesting a conserved model in which the PcG-dependent H3K27me3 mark is inherited by dilution and not by de novo methylation occurring at the time of replication. | | | 22072989
 |