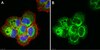
CENP-A nucleosomes localize to transcription factor hotspots and subtelomeric sites in human cancer cells.
Athwal, RK; Walkiewicz, MP; Baek, S; Fu, S; Bui, M; Camps, J; Ried, T; Sung, MH; Dalal, Y
Epigenetics & chromatin
8
2
2015
Mostra il sommario
The histone H3 variant CENP-A is normally tightly regulated to ensure only one centromere exists per chromosome. Native CENP-A is often found overexpressed in human cancer cells and a range of human tumors. Consequently, CENP-A misregulation is thought to contribute to genome instability in human cancers. However, the consequences of such overexpression have not been directly elucidated in human cancer cells.To investigate native CENP-A overexpression, we sought to uncover CENP-A-associated defects in human cells. We confirm that CENP-A is innately overexpressed in several colorectal cancer cell lines. In such cells, we report that a subset of structurally distinct CENP-A-containing nucleosomes associate with canonical histone H3, and with the transcription-coupled chaperones ATRX and DAXX. Furthermore, such hybrid CENP-A nucleosomes localize to DNase I hypersensitive and transcription factor binding sites, including at promoters of genes across the human genome. A distinct class of CENP-A hotspots also accumulates at subtelomeric chromosomal locations, including at the 8q24/Myc region long-associated with genomic instability. We show this 8q24 accumulation of CENP-A can also be seen in early stage primary colorectal tumors.Our data demonstrate that excess CENP-A accumulates at noncentromeric locations in the human cancer genome. These findings suggest that ectopic CENP-A nucleosomes could alter the state of the chromatin fiber, potentially impacting gene regulation and chromosome fragility. | | | 25788983
 |
Nucleolar organization, ribosomal DNA array stability, and acrocentric chromosome integrity are linked to telomere function.
Stimpson, KM; Sullivan, LL; Kuo, ME; Sullivan, BA
PloS one
9
e92432
2014
Mostra il sommario
The short arms of the ten acrocentric human chromosomes share several repetitive DNAs, including ribosomal RNA genes (rDNA). The rDNA arrays correspond to nucleolar organizing regions that coalesce each cell cycle to form the nucleolus. Telomere disruption by expressing a mutant version of telomere binding protein TRF2 (dnTRF2) causes non-random acrocentric fusions, as well as large-scale nucleolar defects. The mechanisms responsible for acrocentric chromosome sensitivity to dysfunctional telomeres are unclear. In this study, we show that TRF2 normally associates with the nucleolus and rDNA. However, when telomeres are crippled by dnTRF2 or RNAi knockdown of TRF2, gross nucleolar and chromosomal changes occur. We used the controllable dnTRF2 system to precisely dissect the timing and progression of nucleolar and chromosomal instability induced by telomere dysfunction, demonstrating that nucleolar changes precede the DNA damage and morphological changes that occur at acrocentric short arms. The rDNA repeat arrays on the short arms decondense, and are coated by RNA polymerase I transcription binding factor UBF, physically linking acrocentrics to one another as they become fusogenic. These results highlight the importance of telomere function in nucleolar stability and structural integrity of acrocentric chromosomes, particularly the rDNA arrays. Telomeric stress is widely accepted to cause DNA damage at chromosome ends, but our findings suggest that it also disrupts chromosome structure beyond the telomere region, specifically within the rDNA arrays located on acrocentric chromosomes. These results have relevance for Robertsonian translocation formation in humans and mechanisms by which acrocentric-acrocentric fusions are promoted by DNA damage and repair. | | | 24662969
 |
Cell-cycle-dependent structural transitions in the human CENP-A nucleosome in vivo.
Bui, M; Dimitriadis, EK; Hoischen, C; An, E; Quénet, D; Giebe, S; Nita-Lazar, A; Diekmann, S; Dalal, Y
Cell
150
317-26
2011
Mostra il sommario
In eukaryotes, DNA is packaged into chromatin by canonical histone proteins. The specialized histone H3 variant CENP-A provides an epigenetic and structural basis for chromosome segregation by replacing H3 at centromeres. Unlike exclusively octameric canonical H3 nucleosomes, CENP-A nucleosomes have been shown to exist as octamers, hexamers, and tetramers. An intriguing possibility reconciling these observations is that CENP-A nucleosomes cycle between octamers and tetramers in vivo. We tested this hypothesis by tracking CENP-A nucleosomal components, structure, chromatin folding, and covalent modifications across the human cell cycle. We report that CENP-A nucleosomes alter from tetramers to octamers before replication and revert to tetramers after replication. These structural transitions are accompanied by reversible chaperone binding, chromatin fiber folding changes, and previously undescribed modifications within the histone fold domains of CENP-A and H4. Our results reveal a cyclical nature to CENP-A nucleosome structure and have implications for the maintenance of epigenetic memory after centromere replication. | | | 22817894
 |
Essential role of microphthalmia transcription factor for DNA replication, mitosis and genomic stability in melanoma.
Strub T, Giuliano S, Ye T, Bonet C, Keime C, Kobi D, Le Gras S, Cormont M, Ballotti R, Bertolotto C, Davidson I
Oncogene
2010
Mostra il sommario
Malignant melanoma is an aggressive cancer known for its notorious resistance to most current therapies. The basic helix-loop-helix microphthalmia transcription factor (MITF) is the master regulator determining the identity and properties of the melanocyte lineage, and is regarded as a lineage-specific \'oncogene\' that has a critical role in the pathogenesis of melanoma. MITF promotes melanoma cell proliferation, whereas sustained supression of MITF expression leads to senescence. By combining chromatin immunoprecipitation coupled to high throughput sequencing (ChIP-seq) and RNA sequencing analyses, we show that MITF directly regulates a set of genes required for DNA replication, repair and mitosis. Our results reveal how loss of MITF regulates mitotic fidelity, and through defective replication and repair induces DNA damage, ultimately ending in cellular senescence. These findings reveal a lineage-specific control of DNA replication and mitosis by MITF, providing new avenues for therapeutic intervention in melanoma. The identification of MITF-binding sites and gene-regulatory networks establish a framework for understanding oncogenic basic helix-loop-helix factors such as N-myc or TFE3 in other cancers.Oncogene advance online publication, 24 January 2011; doi:10.1038/onc.2010.612. | | | 21258399
 |
Uracil DNA N-glycosylase promotes assembly of human centromere protein A.
Zeitlin, SG; Chapados, BR; Baker, NM; Tai, C; Slupphaug, G; Wang, JY
PloS one
6
e17151
2010
Mostra il sommario
Uracil is removed from DNA by the conserved enzyme uracil DNA N-glycosylase (UNG). Previously, we observed that inhibiting UNG in Xenopus egg extracts blocked assembly of CENP-A, a histone H3 variant. CENP-A is an essential protein in all species, since it is required for chromosome segregation during mitosis. Thus, the implication of UNG in CENP-A assembly implies that UNG would also be essential, but UNG mutants lacking catalytic activity are viable in all species. In this paper, we present evidence that UNG2 colocalizes with CENP-A and H2AX phosphorylation at centromeres in normally cycling cells. Reduction of UNG2 in human cells blocks CENP-A assembly, and results in reduced cell proliferation, associated with increased frequencies of mitotic abnormalities and rapid cell death. Overexpression of UNG2 induces high levels of CENP-A assembly in human cells. Using a multiphoton laser approach, we demonstrate that UNG2 is rapidly recruited to sites of DNA damage. Taken together, our data are consistent with a model in which the N-terminus of UNG2 interacts with the active site of the enzyme and with chromatin. | Immunofluorescence | | 21399697
 |
Histone modifications within the human X centromere region.
Mravinac, B; Sullivan, LL; Reeves, JW; Yan, CM; Kopf, KS; Farr, CJ; Schueler, MG; Sullivan, BA
PloS one
4
e6602
2009
Mostra il sommario
Human centromeres are multi-megabase regions of highly ordered arrays of alpha satellite DNA that are separated from chromosome arms by unordered alpha satellite monomers and other repetitive elements. Complexities in assembling such large repetitive regions have limited detailed studies of centromeric chromatin organization. However, a genomic map of the human X centromere has provided new opportunities to explore genomic architecture of a complex locus. We used ChIP to examine the distribution of modified histones within centromere regions of multiple X chromosomes. Methylation of H3 at lysine 4 coincided with DXZ1 higher order alpha satellite, the site of CENP-A localization. Heterochromatic histone modifications were distributed across the 400-500 kb pericentromeric regions. The large arrays of alpha satellite and gamma satellite DNA were enriched for both euchromatic and heterochromatic modifications, implying that some pericentromeric repeats have multiple chromatin characteristics. Partial truncation of the X centromere resulted in reduction in the size of the CENP-A/Cenp-A domain and increased heterochromatic modifications in the flanking pericentromere. Although the deletion removed approximately 1/3 of centromeric DNA, the ratio of CENP-A to alpha satellite array size was maintained in the same proportion, suggesting that a limited, but defined linear region of the centromeric DNA is necessary for kinetochore assembly. Our results indicate that the human X centromere contains multiple types of chromatin, is organized similarly to smaller eukaryotic centromeres, and responds to structural changes by expanding or contracting domains. Testo completo dell'articolo | | Human | 19672304
 |
GNL3L stabilizes the TRF1 complex and promotes mitotic transition.
Zhu, Qubo, et al.
J. Cell Biol., 185: 827-39 (2009)
2009
Mostra il sommario
Telomeric repeat binding factor 1 (TRF1) is a component of the multiprotein complex "shelterin," which organizes the telomere into a high-order structure. TRF1 knockout embryos suffer from severe growth defects without apparent telomere dysfunction, suggesting an obligatory role for TRF1 in cell cycle control. To date, the mechanism regulating the mitotic increase in TRF1 protein expression and its function in mitosis remains unclear. Here, we identify guanine nucleotide-binding protein-like 3 (GNL3L), a GTP-binding protein most similar to nucleostemin, as a novel TRF1-interacting protein in vivo. GNL3L binds TRF1 in the nucleoplasm and is capable of promoting the homodimerization and telomeric association of TRF1, preventing promyelocytic leukemia body recruitment of telomere-bound TRF1, and stabilizing TRF1 protein by inhibiting its ubiquitylation and binding to FBX4, an E3 ubiquitin ligase for TRF1. Most importantly, the TRF1 protein-stabilizing activity of GNL3L mediates the mitotic increase of TRF1 protein and promotes the metaphase-to-anaphase transition. This work reveals novel aspects of TRF1 modulation by GNL3L. | Immunocytochemistry | Human | 19487455
 |